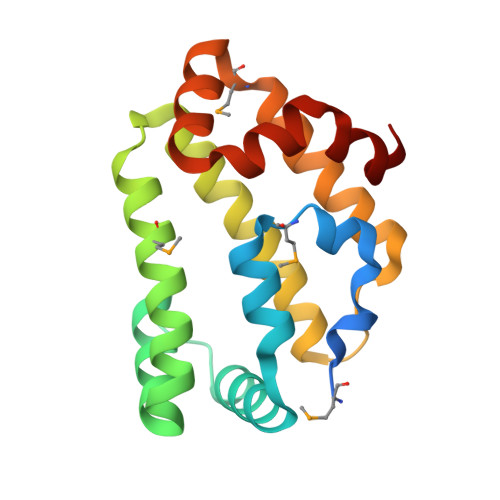
Diversity in the structures and ligand-binding sites of nematode fatty acid and retinol-binding proteins revealed by Na-FAR-1 from Necator americanus.
Rey-Burusco, M.F., Ibanez-Shimabukuro, M., Gabrielsen, M., Franchini, G.R., Roe, A.J., Griffiths, K., Zhan, B., Cooper, A., Kennedy, M.W., Corsico, B., Smith, B.O.(2015) Biochem J 471: 403-414
- PubMed: 26318523
- DOI: https://doi.org/10.1042/BJ20150068
- Primary Citation of Related Structures:
4UET, 4XCP - PubMed Abstract:
Fatty acid and retinol-binding proteins (FARs) comprise a family of unusual α-helix rich lipid-binding proteins found exclusively in nematodes. They are secreted into host tissues by parasites of plants, animals and humans. The structure of a FAR protein from the free-living nematode Caenorhabditis elegans is available, but this protein [C. elegans FAR-7 (Ce-FAR-7)] is from a subfamily of FARs that does not appear to be important at the host/parasite interface. We have therefore examined [Necator americanus FAR-1 (Na-FAR-1)] from the blood-feeding intestinal parasite of humans, N. americanus. The 3D structure of Na-FAR-1 in its ligand-free and ligand-bound forms, determined by NMR (nuclear magnetic resonance) spectroscopy and X-ray crystallography respectively, reveals an α-helical fold similar to Ce-FAR-7, but Na-FAR-1 possesses a larger and more complex internal ligand-binding cavity and an additional C-terminal α-helix. Titration of apo-Na-FAR-1 with oleic acid, analysed by NMR chemical shift perturbation, reveals that at least four distinct protein-ligand complexes can be formed. Na-FAR-1 and possibly other FARs may have a wider repertoire for hydrophobic ligand binding, as confirmed in the present study by our finding that a range of neutral and polar lipids co-purify with the bacterially expressed recombinant protein. Finally, we show by immunohistochemistry that Na-FAR-1 is present in adult worms with a tissue distribution indicative of possible roles in nutrient acquisition by the parasite and in reproduction in the male.
- Instituto de Investigaciones Bioquímicas de La Plata, CONICET-UNLP, Facultad de Ciencias Médicas, calles 60 y 120, 1900-La Plata, Argentina.
Organizational Affiliation: