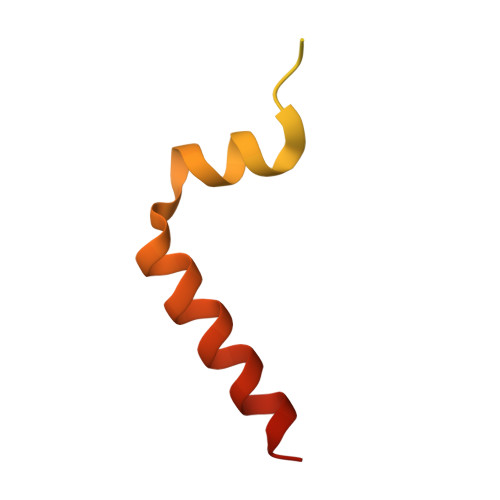
Structural, functional, and genetic analyses of the actinobacterial transcription factor RbpA.
Hubin, E.A., Tabib-Salazar, A., Humphrey, L.J., Flack, J.E., Olinares, P.D., Darst, S.A., Campbell, E.A., Paget, M.S.(2015) Proc Natl Acad Sci U S A 112: 7171-7176
- PubMed: 26040003 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1504942112
- Primary Citation Related Structures:
4X8K - PubMed Abstract:
Gene expression is highly regulated at the step of transcription initiation, and transcription activators play a critical role in this process. RbpA, an actinobacterial transcription activator that is essential in Mycobacterium tuberculosis (Mtb), binds selectively to group 1 and certain group 2 σ-factors. To delineate the molecular mechanism of RbpA, we show that the Mtb RbpA σ-interacting domain (SID) and basic linker are sufficient for transcription activation. We also present the crystal structure of the Mtb RbpA-SID in complex with domain 2 of the housekeeping σ-factor, σ(A). The structure explains the basis of σ-selectivity by RbpA, showing that RbpA interacts with conserved regions of σ(A) as well as the nonconserved region (NCR), which is present only in housekeeping σ-factors. Thus, the structure is the first, to our knowledge, to show a protein interacting with the NCR of a σ-factor. We confirm the basis of selectivity and the observed interactions using mutagenesis and functional studies. In addition, the structure allows for a model of the RbpA-SID in the context of a transcription initiation complex. Unexpectedly, the structural modeling suggests that RbpA contacts the promoter DNA, and we present in vivo and in vitro studies supporting this finding. Our combined data lead to a better understanding of the mechanism of RbpA function as a transcription activator.
- Laboratory of Molecular Biophysics, The Rockefeller University, New York, NY 10065;
Organizational Affiliation: