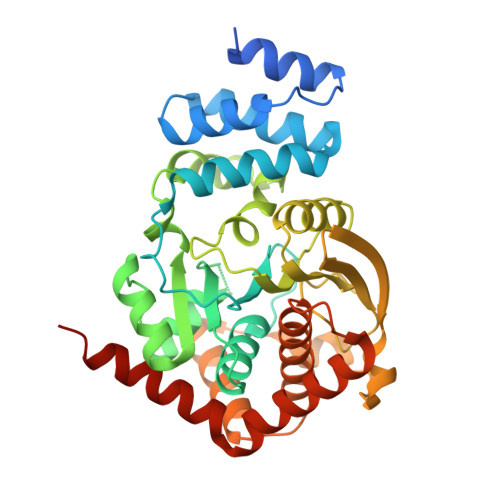
Structures of Mycobacterium tuberculosis Anthranilate Phosphoribosyltransferase Variants Reveal the Conformational Changes That Facilitate Delivery of the Substrate to the Active Site.
Cookson, T.V., Evans, G.L., Castell, A., Baker, E.N., Lott, J.S., Parker, E.J.(2015) Biochemistry 54: 6082-6092
- PubMed: 26356348
- DOI: https://doi.org/10.1021/acs.biochem.5b00612
- Primary Citation of Related Structures:
4X58, 4X59, 4X5A, 4X5B, 4X5C, 4X5D, 4X5E - PubMed Abstract:
Anthranilate phosphoribosyltransferase (AnPRT) is essential for the biosynthesis of tryptophan in Mycobacterium tuberculosis (Mtb). This enzyme catalyzes the second committed step in tryptophan biosynthesis, the Mg²⁺-dependent reaction between 5'-phosphoribosyl-1'-pyrophosphate (PRPP) and anthranilate. The roles of residues predicted to be involved in anthranilate binding have been tested by the analysis of six Mtb-AnPRT variant proteins. Kinetic analysis showed that five of six variants were active and identified the conserved residue R193 as being crucial for both anthranilate binding and catalytic function. Crystal structures of these Mtb-AnPRT variants reveal the ability of anthranilate to bind in three sites along an extended anthranilate tunnel and expose the role of the mobile β2-α6 loop in facilitating the enzyme's sequential reaction mechanism. The β2-α6 loop moves sequentially between a "folded" conformation, partially occluding the anthranilate tunnel, via an "open" position to a "closed" conformation, which supports PRPP binding and allows anthranilate access via the tunnel to the active site. The return of the β2-α6 loop to the "folded" conformation completes the catalytic cycle, concordantly allowing the active site to eject the product PRA and rebind anthranilate at the opening of the anthranilate tunnel for subsequent reactions. Multiple anthranilate molecules blocking the anthranilate tunnel prevent the β2-α6 loop from undergoing the conformational changes required for catalysis, thus accounting for the unusual substrate inhibition of this enzyme.
- Maurice Wilkins Centre for Molecular Biodiscovery, Biomolecular Interaction Centre, and Department of Chemistry, University of Canterbury , 20 Kirkwood Avenue, Christchurch 8140, New Zealand.
Organizational Affiliation: