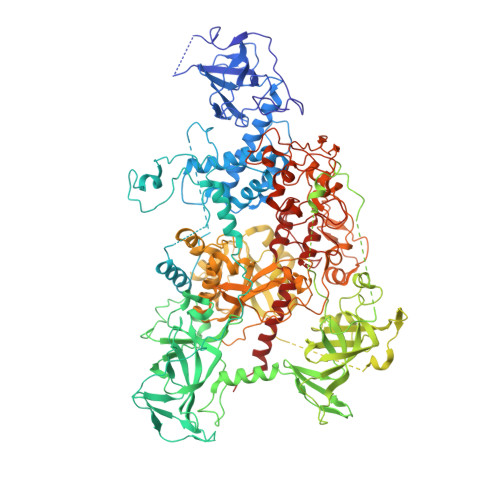
Crystal Structure of Human DNA Methyltransferase 1.
Zhang, Z.M., Liu, S., Lin, K., Luo, Y., Perry, J.J., Wang, Y., Song, J.(2015) J Mol Biology 427: 2520-2531
- PubMed: 26070743
- DOI: https://doi.org/10.1016/j.jmb.2015.06.001
- Primary Citation Related Structures:
4WXX - PubMed Abstract:
DNMT1 (DNA methyltransferase 1) is responsible for propagating the DNA methylation patterns during DNA replication. DNMT1 contains, in addition to a C-terminal methyltransferase domain, a large N-terminal regulatory region that is composed of an RFTS (replication foci targeting sequence) domain, a CXXC zinc finger domain and a pair of BAH (bromo adjacent homology) domains. The regulatory domains of DNMT1 mediate a network of protein-protein and protein-DNA interactions to control the recruitment and enzymatic activity of DNMT1. Here we report the crystal structure of human DNMT1 with all the structural domains (hDNMT1, residues 351-1600) in complex with S-adenosyl-l-homocysteine at 2.62Å resolution. The RFTS domain directly associates with the methyltransferase domain, thereby inhibiting the substrate binding of hDNMT1. Through structural analysis, mutational, biochemical and enzymatic studies, we further identify that a linker sequence between the CXXC and BAH1 domains, aside from its role in the CXXC domain-mediated DNMT1 autoinhibition, serves as an important regulatory element in the RFTS domain-mediated autoinhibition. In comparison with the previously determined structure of mouse DNMT1, this study also reveals a number of distinct structural features that may underlie subtle functional diversity observed for the two orthologues. In addition, this structure provides a framework for understanding the functional consequence of disease-related hDNMT1 mutations.
- Department of Biochemistry, University of California, Riverside, CA 92521, USA.
Organizational Affiliation: