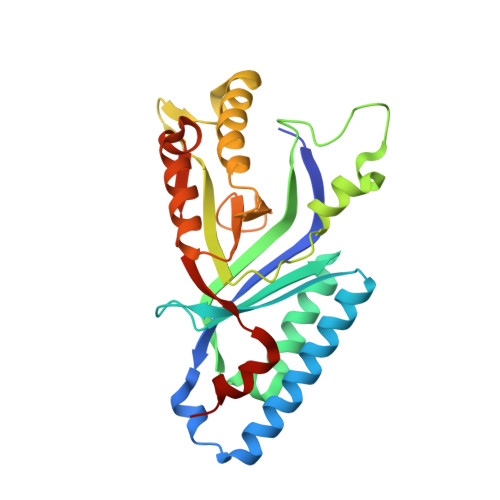
Structure and heme-binding properties of HemQ (chlorite dismutase-like protein) from Listeria monocytogenes.
Hofbauer, S., Hagmuller, A., Schaffner, I., Mlynek, G., Krutzler, M., Stadlmayr, G., Pirker, K.F., Obinger, C., Daims, H., Djinovic-Carugo, K., Furtmuller, P.G.(2015) Arch Biochem Biophys 574: 36-48
- PubMed: 25602700
- DOI: https://doi.org/10.1016/j.abb.2015.01.010
- Primary Citation Related Structures:
4WWS - PubMed Abstract:
Chlorite dismutase-like proteins are structurally closely related to functional chlorite dismutases which are heme b-dependent oxidoreductases capable of reducing chlorite to chloride with simultaneous production of dioxygen. Chlorite dismutase-like proteins are incapable of performing this reaction and their biological role is still under discussion. Recently, members of this large protein family were shown to be involved in heme biosynthesis in Gram-positive bacteria, and thus the protein was renamed HemQ in these organisms. In the present work the structural and heme binding properties of the chlorite dismutase-like protein from the Gram-positive pathogen Listeria monocytogenes (LmCld) were analyzed in order to evaluate its potential role as a regulatory heme sensing protein. The homopentameric crystal structure (2.0Å) shows high similarity to chlorite-degrading chlorite dismutases with an important difference in the structure of the putative substrate and heme entrance channel. In solution LmCld is a stable hexamer able to bind the low-spin ligand cyanide. Heme binding is reversible with KD-values determined to be 7.2μM (circular dichroism spectroscopy) and 16.8μM (isothermal titration calorimetry) at pH 7.0. Both acidic and alkaline conditions promote heme release. Presented biochemical and structural data reveal that the chlorite dismutase-like protein from L. monocytogenes could act as a potential regulatory heme sensing and storage protein within heme biosynthesis.
- Department of Chemistry, Division of Biochemistry, BOKU - University of Natural Resources and Life Sciences, Muthgasse 18, A-1190 Vienna, Austria; Department for Structural and Computational Biology, Max F. Perutz Laboratories, University of Vienna, A-1030 Vienna, Austria.
Organizational Affiliation: