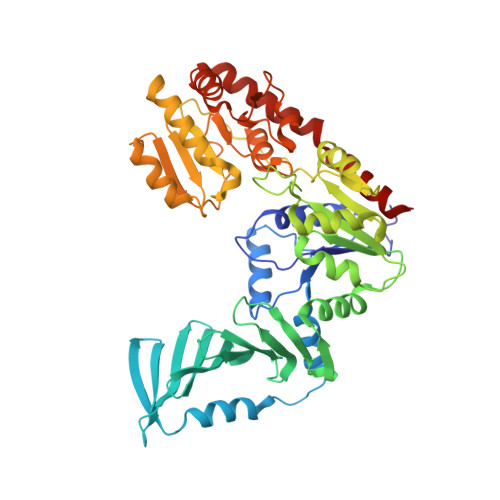
Structural and Enzymatic Analysis of TarM Glycosyltransferase from Staphylococcus aureus Reveals an Oligomeric Protein Specific for the Glycosylation of Wall Teichoic Acid.
Koc, C., Gerlach, D., Beck, S., Peschel, A., Xia, G., Stehle, T.(2015) J Biological Chem 290: 9874-9885
- PubMed: 25697358 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.619924
- Primary Citation Related Structures:
4WAC, 4WAD - PubMed Abstract:
Anionic glycopolymers known as wall teichoic acids (WTAs) functionalize the peptidoglycan layers of many Gram-positive bacteria. WTAs play central roles in many fundamental aspects of bacterial physiology, and they are important determinants of pathogenesis and antibiotic resistance. A number of enzymes that glycosylate WTA in Staphylococcus aureus have recently been identified. Among these is the glycosyltransferase TarM, a component of the WTA de novo biosynthesis pathway. TarM performs the synthesis of α-O-N-acetylglycosylated poly-5'-phosphoribitol in the WTA structure. We have solved the crystal structure of TarM at 2.4 Å resolution, and we have also determined a structure of the enzyme in complex with its substrate UDP-GlcNAc at 2.8 Å resolution. The protein assembles into a propeller-like homotrimer in which each blade contains a GT-B-type glycosyltransferase domain with a typical Rossmann fold. The enzymatic reaction retains the stereochemistry of the anomeric center of the transferred GlcNAc-moiety on the polyribitol backbone. TarM assembles into a trimer using a novel trimerization domain, here termed the HUB domain. Structure-guided mutagenesis experiments of TarM identify residues critical for enzyme activity, assign a putative role for the HUB in TarM function, and allow us to propose a likely reaction mechanism.
- From the Interfaculty Institute of Biochemistry, University of Tübingen, 72076 Tübingen, Germany.
Organizational Affiliation: