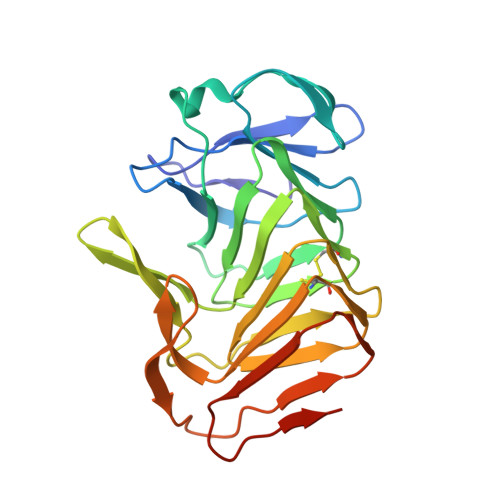
Proteolysis of truncated hemolysin A yields a stable dimerization interface.
Novak, W.R., Bhattacharyya, B., Grilley, D.P., Weaver, T.M.(2017) Acta Crystallogr F Struct Biol Commun 73: 138-145
- PubMed: 28291749
- DOI: https://doi.org/10.1107/S2053230X17002102
- Primary Citation of Related Structures:
4W8Q, 5KEH, 5KF3, 5KKD, 5SZ8 - PubMed Abstract:
Wild-type and variant forms of HpmA265 (truncated hemolysin A) from Proteus mirabilis reveal a right-handed, parallel β-helix capped and flanked by segments of antiparallel β-strands. The low-salt crystal structures form a dimeric structure via the implementation of on-edge main-chain hydrogen bonds donated by residues 243-263 of adjacent monomers. Surprisingly, in the high-salt structures of two variants, Y134A and Q125A-Y134A, a new dimeric interface is formed via main-chain hydrogen bonds donated by residues 203-215 of adjacent monomers, and a previously unobserved tetramer is formed. In addition, an eight-stranded antiparallel β-sheet is formed from the flap regions of crystallographically related monomers in the high-salt structures. This new interface is possible owing to additional proteolysis of these variants after Tyr240. The interface formed in the high-salt crystal forms of hemolysin A variants may mimic the on-edge β-strand positioning used in template-assisted hemolytic activity.
- Department of Chemistry, Wabash College, 301 West Wabash Avenue, Crawfordsville, IN 47933, USA.
Organizational Affiliation: