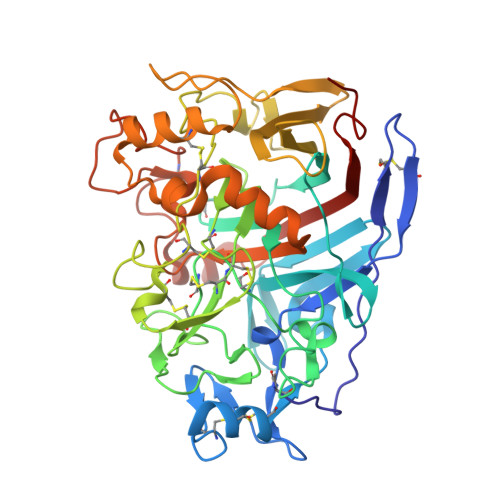
The Three-Dimensional Structure of the Cellobiohydrolase Cel7A from Aspergillus Fumigatus at 1.5 A Resolution
Moroz, O.V., Maranta, M., Shaghasi, T., Harris, P.V., Wilson, K.S., Davies, G.J.(2015) Acta Crystallogr Sect F Struct Biol Cryst Commun 71: 114
- PubMed: 25615982
- DOI: https://doi.org/10.1107/S2053230X14027307
- Primary Citation Related Structures:
4V1Z, 4V20 - PubMed Abstract:
The enzymatic degradation of plant cell-wall cellulose is central to many industrial processes, including second-generation biofuel production. Key players in this deconstruction are the fungal cellobiohydrolases (CBHs), notably those from family GH7 of the carbohydrate-active enzymes (CAZY) database, which are generally known as CBHI enzymes. Here, three-dimensional structures are reported of the Aspergillus fumigatus CBHI Cel7A solved in uncomplexed and disaccharide-bound forms at resolutions of 1.8 and 1.5 Å, respectively. The product complex with a disaccharide in the +1 and +2 subsites adds to the growing three-dimensional insight into this family of industrially relevant biocatalysts.
- Department of Chemistry, University of York, York Structural Biology Laboratory, York YO10 5DD, England.
Organizational Affiliation: