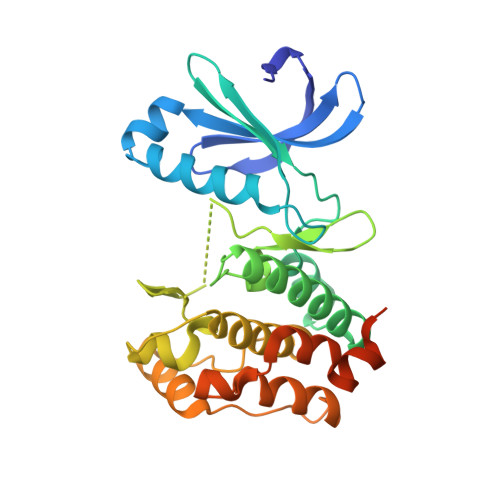
Sar156497 an Exquisitely Selective Inhibitor of Aurora Kinases.
Carry, J.C., Clerc, F.F., Minoux, H., Schio, L., Mauger, J., Nair, A., Parmantier, E., Le Moigne, R., Delorme, C., Nicolas, J.P., Krick, A., Abecassis, P.Y., Crocq-Stuerga, V., Pouzieux, S., Delarbre, L., Maignan, S., Bertrand, T., Bjergarde, K., Ma, N., Lachaud, S., Guizani, H., Lebel, R., Doerflinger, G., Monget, S., Perron, S., Gasse, F., Angouillant-Boniface, O., Filoche-Romme, B.J., Murer, M., Gontier, S., Prevost, C., Monteiro, M.L., Combeau, C.(2015) J Med Chem 58: 362
- PubMed: 25369539 Search on PubMed
- DOI: https://doi.org/10.1021/jm501326k
- Primary Citation Related Structures:
4UYN, 4UZD, 4UZH - PubMed Abstract:
The Aurora family of serine/threonine kinases is essential for mitosis. Their crucial role in cell cycle regulation and aberrant expression in a broad range of malignancies have been demonstrated and have prompted intensive search for small molecule Aurora inhibitors. Indeed, over 10 of them have reached the clinic as potential anticancer therapies. We report herein the discovery and optimization of a novel series of tricyclic molecules that has led to SAR156497, an exquisitely selective Aurora A, B, and C inhibitor with in vitro and in vivo efficacy. We also provide insights into its mode of binding to its target proteins, which could explain its selectivity.
- Oncology Drug Discovery, ‡Structure Design Informatics, §Disposition Safety Animal Research, ∥Chemical Development, and ⊥Analytical Sciences, Sanofi , 13 Quai Jules Guesde, 94403 Vitry-sur-Seine, France.
Organizational Affiliation: