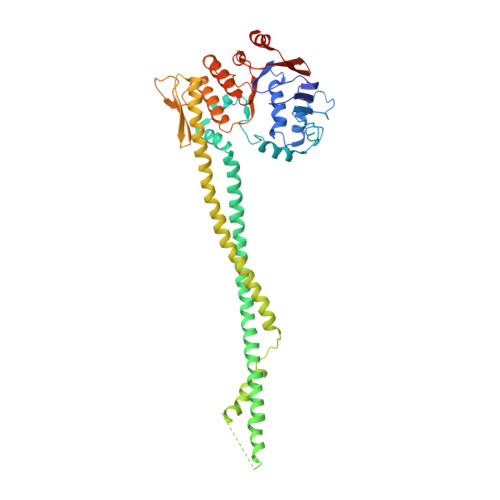
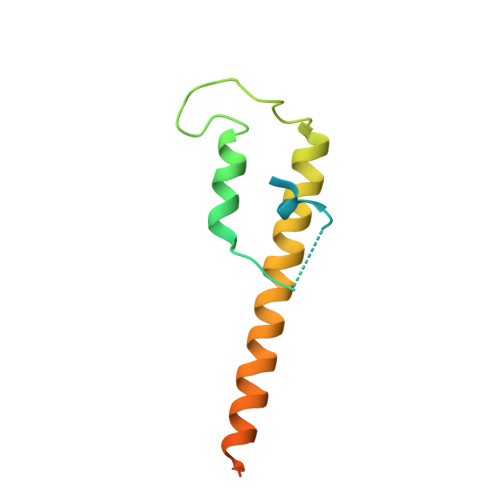
Closing the Cohesin Ring: Structure and Function of its Smc3-Kleisin Interface.
Gligoris, T.G., Scheinost, J.C., Burmann, F., Petela, N., Chan, K., Uluocak, P., Beckouet, F., Gruber, S., Nasmyth, K., Lowe, J.(2014) Science 346: 963
- PubMed: 25414305
- DOI: https://doi.org/10.1126/science.1256917
- Primary Citation Related Structures:
4UX3 - PubMed Abstract:
Through their association with a kleisin subunit (Scc1), cohesin's Smc1 and Smc3 subunits are thought to form tripartite rings that mediate sister chromatid cohesion. Unlike the structure of Smc1/Smc3 and Smc1/Scc1 interfaces, that of Smc3/Scc1 is not known. Disconnection of this interface is thought to release cohesin from chromosomes in a process regulated by acetylation. We show here that the N-terminal domain of yeast Scc1 contains two α helices, forming a four-helix bundle with the coiled coil emerging from Smc3's adenosine triphosphatase head. Mutations affecting this interaction compromise cohesin's association with chromosomes. The interface is far from Smc3 residues, whose acetylation prevents cohesin's dissociation from chromosomes. Cohesin complexes holding chromatids together in vivo do indeed have the configuration of hetero-trimeric rings, and sister DNAs are entrapped within these.
- Department of Biochemistry, University of Oxford, Oxford, OX1 3QU, UK.
Organizational Affiliation: