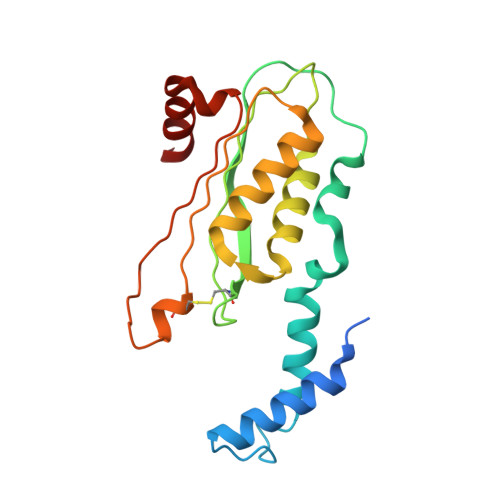
Structure of GTP cyclohydrolase I from Listeria monocytogenes, a potential anti-infective drug target.
Schussler, S., Haase, I., Perbandt, M., Illarionov, B., Siemens, A., Richter, K., Bacher, A., Fischer, M., Grawert, T.(2019) Acta Crystallogr F Struct Biol Commun 75: 586-592
- PubMed: 31475925
- DOI: https://doi.org/10.1107/S2053230X19010902
- Primary Citation Related Structures:
4UQF - PubMed Abstract:
A putative open reading frame encoding GTP cyclohydrolase I from Listeria monocytogenes was expressed in a recombinant Escherichia coli strain. The recombinant protein was purified and was confirmed to convert GTP to dihydroneopterin triphosphate (K m = 53 µM; v max = 180 nmol mg -1 min -1 ). The protein was crystallized from 1.3 M sodium citrate pH 7.3 and the crystal structure was solved at a resolution of 2.4 Å (R free = 0.226) by molecular replacement using human GTP cyclohydrolase I as a template. The protein is a D 5 -symmetric decamer with ten topologically equivalent active sites. Screening a small library of about 9000 compounds afforded several inhibitors with IC 50 values in the low-micromolar range. Several inhibitors had significant selectivity with regard to human GTP cyclohydrolase I. Hence, GTP cyclohydrolase I may be a potential target for novel drugs directed at microbial infections, including listeriosis, a rare disease with high mortality.
- Hamburg School of Food Science, Universität Hamburg, Grindelallee 117, 20146 Hamburg, Germany.
Organizational Affiliation: