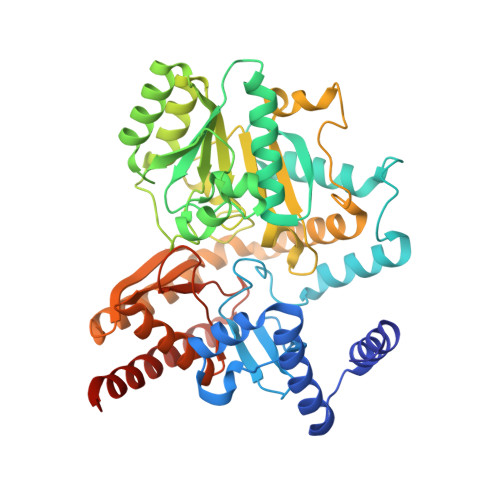
Structure of Putrescine Aminotransferase from Escherichia Coli Provides Insights Into the Substrate Specificity Among Class III Aminotransferases.
Cha, H.J., Jeong, J., Rojviriya, C., Kim, Y.(2014) PLoS One 9: 13212
- PubMed: 25423189 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0113212
- Primary Citation Related Structures:
4UOX, 4UOY - PubMed Abstract:
YgjG is a putrescine aminotransferase enzyme that transfers amino groups from compounds with terminal primary amines to compounds with an aldehyde group using pyridoxal-5'-phosphate (PLP) as a cofactor. Previous biochemical data show that the enzyme prefers primary diamines, such as putrescine, over ornithine as a substrate. To better understand the enzyme's substrate specificity, crystal structures of YgjG from Escherichia coli were determined at 2.3 and 2.1 Å resolutions for the free and putrescine-bound enzymes, respectively. Sequence and structural analyses revealed that YgjG forms a dimer that adopts a class III PLP-dependent aminotransferase fold. A structural comparison between YgjG and other class III aminotransferases revealed that their structures are similar. However, YgjG has an additional N-terminal helical structure that partially contributes to a dimeric interaction with the other subunit via a helix-helix interaction. Interestingly, the YgjG substrate-binding site entrance size and charge distribution are smaller and more hydrophobic than other class III aminotransferases, which suggest that YgjG has a unique substrate binding site that could accommodate primary aliphatic diamine substrates, including putrescine. The YgjG crystal structures provide structural clues to putrescine aminotransferase substrate specificity and binding.
- Pohang Accelerator Laboratory, Pohang University of Science and Technology, Pohang, Korea.
Organizational Affiliation: