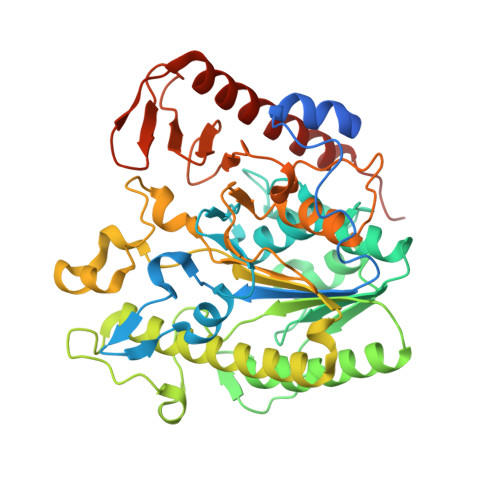
Structural and mechanistic insight into the Listeria monocytogenes two-enzyme lipoteichoic acid synthesis system.
Campeotto, I., Percy, M.G., MacDonald, J.T., Forster, A., Freemont, P.S., Grundling, A.(2014) J Biological Chem 289: 28054-28069
- PubMed: 25128528 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.590570
- Primary Citation Related Structures:
4UOO, 4UOP, 4UOR - PubMed Abstract:
Lipoteichoic acid (LTA) is an important cell wall component required for proper cell growth in many Gram-positive bacteria. In Listeria monocytogenes, two enzymes are required for the synthesis of this polyglycerolphosphate polymer. The LTA primase LtaP(Lm) initiates LTA synthesis by transferring the first glycerolphosphate (GroP) subunit onto the glycolipid anchor and the LTA synthase LtaS(Lm) extends the polymer by the repeated addition of GroP subunits to the tip of the growing chain. Here, we present the crystal structures of the enzymatic domains of LtaP(Lm) and LtaS(Lm). Although the enzymes share the same fold, substantial differences in the cavity of the catalytic site and surface charge distribution contribute to enzyme specialization. The eLtaS(Lm) structure was also determined in complex with GroP revealing a second GroP binding site. Mutational analysis confirmed an essential function for this binding site and allowed us to propose a model for the binding of the growing chain.
- From the Section of Microbiology and MRC Centre for Molecular Bacteriology and Infection, and.
Organizational Affiliation: