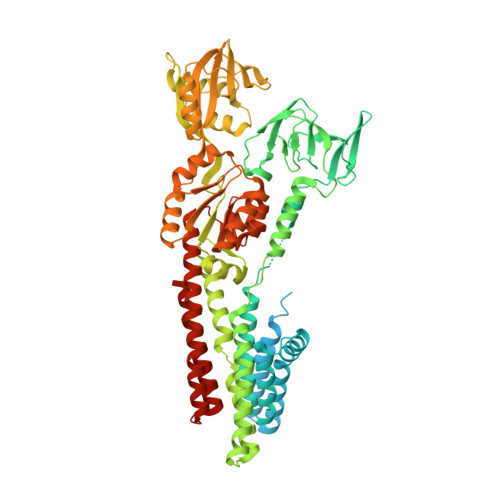
Structure and Mechanism of Zn(2+)-Transporting P-Type Atpases.
Wang, K., Sitsel, O., Meloni, G., Autzen, H.E., Andersson, M., Klymchuk, T., Nielsen, A.M., Rees, D.C., Nissen, P., Gourdon, P.(2014) Nature 514: 518
- PubMed: 25132545 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature13618
- Primary Citation Related Structures:
4UMV, 4UMW - PubMed Abstract:
Zinc is an essential micronutrient for all living organisms. It is required for signalling and proper functioning of a range of proteins involved in, for example, DNA binding and enzymatic catalysis. In prokaryotes and photosynthetic eukaryotes, Zn(2+)-transporting P-type ATPases of class IB (ZntA) are crucial for cellular redistribution and detoxification of Zn(2+) and related elements. Here we present crystal structures representing the phosphoenzyme ground state (E2P) and a dephosphorylation intermediate (E2·Pi) of ZntA from Shigella sonnei, determined at 3.2 Å and 2.7 Å resolution, respectively. The structures reveal a similar fold to Cu(+)-ATPases, with an amphipathic helix at the membrane interface. A conserved electronegative funnel connects this region to the intramembranous high-affinity ion-binding site and may promote specific uptake of cellular Zn(2+) ions by the transporter. The E2P structure displays a wide extracellular release pathway reaching the invariant residues at the high-affinity site, including C392, C394 and D714. The pathway closes in the E2·Pi state, in which D714 interacts with the conserved residue K693, which possibly stimulates Zn(2+) release as a built-in counter ion, as has been proposed for H(+)-ATPases. Indeed, transport studies in liposomes provide experimental support for ZntA activity without counter transport. These findings suggest a mechanistic link between PIB-type Zn(2+)-ATPases and PIII-type H(+)-ATPases and at the same time show structural features of the extracellular release pathway that resemble PII-type ATPases such as the sarcoplasmic/endoplasmic reticulum Ca(2+)-ATPase (SERCA) and Na(+), K(+)-ATPase. These findings considerably increase our understanding of zinc transport in cells and represent new possibilities for biotechnology and biomedicine.
- 1] Centre for Membrane Pumps in Cells and Disease (PUMPkin), Danish National Research Foundation, Aarhus University, Department of Molecular Biology and Genetics, Gustav Wieds Vej 10C, DK-8000 Aarhus C, Denmark [2] Department of Biomedical Sciences, University of Copenhagen, Blegdamsvej 3B, DK-2200 Copenhagen, Denmark (K.W. and P.G.); Department of Experimental Medical Science, Lund University, Sölvegatan 19, SE-221 84 Lund, Sweden (P.G.). [3].
Organizational Affiliation: