Negishi Cross-Coupling Enabled Synthesis of Novel Nad(+)-Dependent DNA Ligase Inhibitors and Sar Development.
Murphy-Benenato, K.E., Gingipalli, L., Boriack-Sjodin, P.A., Martinez-Botella, G., Carcanague, D., Eyermann, C.J., Gowravaram, M., Harang, J., Hale, M.R., Ioannidis, G., Jahic, H., Johnstone, M., Kutschke, A., Laganas, V.A., Loch, J.T., Miller, M.D., Oguto, H., Patel, S.J.(2015) Bioorg Med Chem Lett 25: 5172
- PubMed: 26463129
- DOI: https://doi.org/10.1016/j.bmcl.2015.09.075
- Primary Citation of Related Structures:
4UFZ - PubMed Abstract:
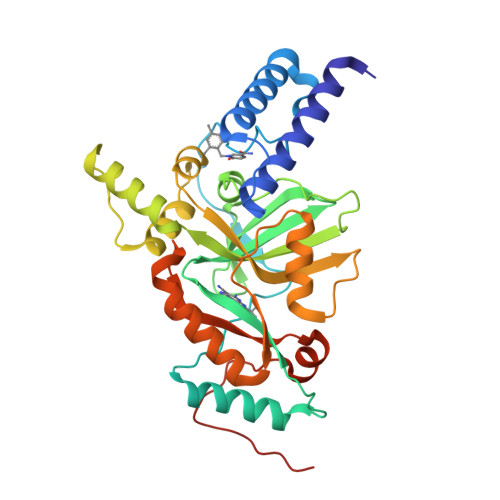
Two novel compounds, pyridopyrimidines (1) and naphthyridines (2) were identified as potent inhibitors of bacterial NAD(+)-dependent DNA ligase (Lig) A in a fragment screening. SAR was guided by molecular modeling and X-ray crystallography. It was observed that the diaminonitrile pharmacophore made a key interaction with the ligase enzyme, specifically residues Glu114, Lys291, and Leu117. Synthetic challenges limited opportunities for diversification of the naphthyridine core, therefore most of the SAR was focused on a pyridopyrimidine scaffold. The initial diversification at R(1) improved both enzyme and cell potency. Further SAR developed at the R(2) position using the Negishi cross-coupling reaction provided several compounds, among these compounds 22g showed good enzyme potency and cellular potency.
Organizational Affiliation:
Infection Innovative Medicines Unit, AstraZeneca R&D Boston, 35 Gatehouse Drive, Waltham, MA 02451, USA.