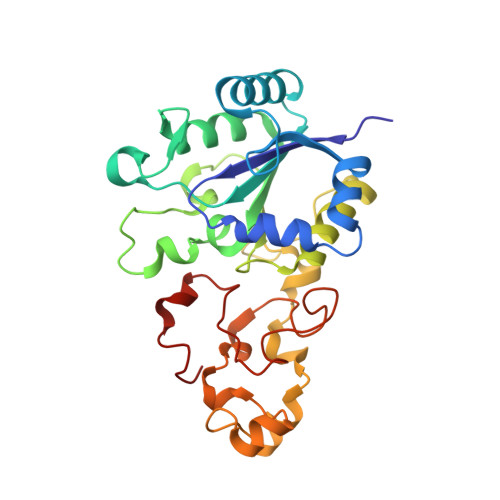
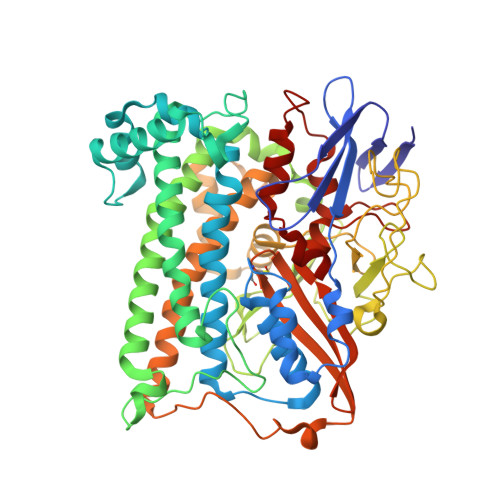
Mechanism of Hydrogen Activation by [Nife] Hydrogenases.
Evans, R.M., Brooke, E.J., Wehlin, S.A., Nomerotskaia, E., Sargent, F., Carr, S.B., Phillips, S.E., Armstrong, F.A.(2016) Nat Chem Biol 12: 46
- PubMed: 26619250
- DOI: https://doi.org/10.1038/nchembio.1976
- Primary Citation Related Structures:
4UE3, 5A4F, 5A4I, 5A4M, 5ADU - PubMed Abstract:
The active site of [NiFe] hydrogenases contains a strictly conserved arginine that suspends a guanidine nitrogen atom <4.5 Å above the nickel and iron atoms. The guanidine headgroup interacts with the side chains of two conserved aspartic acid residues to complete an outer-shell canopy that has thus far proved intractable to investigation by site-directed mutagenesis. Using hydrogenase-1 from Escherichia coli, the strictly conserved residues R509 and D574 have been replaced by lysine (R509K) and asparagine (D574N) and the highly conserved D118 has been replaced by alanine (D118A) or asparagine (D118N/D574N). Each enzyme variant is stable, and their [(RS)2Niμ(SR)2Fe(CO)(CN)2] inner coordination shells are virtually unchanged. The R509K variant had >100-fold lower activity than native enzyme. Conversely, the variants D574N, D118A and D118N/D574N, in which the position of the guanidine headgroup is retained, showed 83%, 26% and 20% activity, respectively. The special kinetic requirement for R509 implicates the suspended guanidine group as the general base in H2 activation by [NiFe] hydrogenases.
- Department of Chemistry, University of Oxford, Oxford, UK.
Organizational Affiliation: