Unraveling the Specific Regulation of the Mbd Central Pathway for the Anaerobic Degradation of 3-Methylbenzoate.
Juarez, J.F., Liu, H., Zamarro, M.T., Mcmahon, S., Liu, H., Naismith, J.H., Eberlein, C., Boll, M., Carmona, M., Diaz, E.(2015) J Biological Chem 290: 12165
- PubMed: 25795774
- DOI: https://doi.org/10.1074/jbc.M115.637074
- Primary Citation Related Structures:
4UDS - PubMed Abstract:
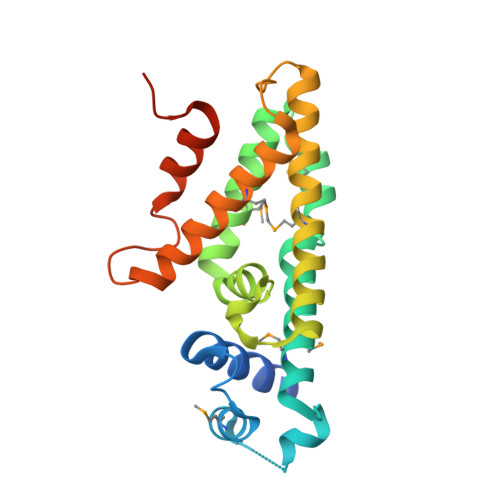
The mbd cluster encodes the anaerobic degradation of 3-methylbenzoate in the β-proteobacterium Azoarcus sp. CIB. The specific transcriptional regulation circuit that controls the expression of the mbd genes was investigated. The PO, PB 1, and P3 R promoters responsible for the expression of the mbd genes, their cognate MbdR transcriptional repressor, as well as the MbdR operator regions (ATACN10GTAT) have been characterized. The three-dimensional structure of MbdR has been solved revealing a conformation similar to that of other TetR family transcriptional regulators. The first intermediate of the catabolic pathway, i.e. 3-methylbenzoyl-CoA, was shown to act as the inducer molecule. An additional MbdR-dependent promoter, PA, which contributes to the expression of the CoA ligase that activates 3-methylbenzoate to 3-methylbenzoyl-CoA, was shown to be necessary for an efficient induction of the mbd genes. Our results suggest that the mbd cluster recruited a regulatory system based on the MbdR regulator and its target promoters to evolve a distinct central catabolic pathway that is only expressed for the anaerobic degradation of aromatic compounds that generate 3-methylbenzoyl-CoA as the central metabolite. All these results highlight the importance of the regulatory systems in the evolution and adaptation of bacteria to the anaerobic degradation of aromatic compounds.
- From the Department of Environmental Biology, Centro de Investigaciones Biológicas-Consejo Superior de Investigaciones Científicas, Ramiro de Maeztu 9, 28040 Madrid, Spain.
Organizational Affiliation: