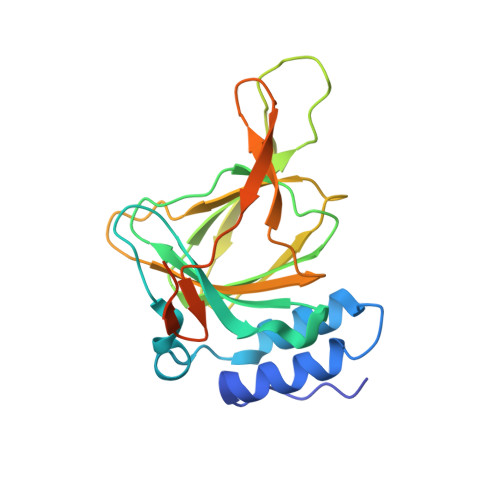
The Cys-Tyr Cross-Link of Cysteine Dioxygenase Changes the Optimal pH of the Reaction without a Structural Change.
Davies, C.G., Fellner, M., Tchesnokov, E.P., Wilbanks, S.M., Jameson, G.N.(2014) Biochemistry 53: 7961-7968
- PubMed: 25390690 Search on PubMed
- DOI: https://doi.org/10.1021/bi501277a
- Primary Citation Related Structures:
4UBG - PubMed Abstract:
Cysteine dioxygenase (CDO) is a non-heme monoiron enzyme with an unusual posttranslational modification in the proximity of the ferrous iron active site. This modification, a cysteine to tyrosine thioether bond, cross-links two β-strands of the β-barrel. We have investigated its role in catalysis through a combined crystallographic and kinetic approach. The C93G variant lacks the cross-link and shows little change in structure from that of the wild type, suggesting that the cross-link does not stabilize an otherwise unfavorable conformation. A pH-dependent kinetic study shows that both cross-linked and un-cross-linked CDO are active but the optimal pH decreases with the presence of the cross-link. This result reflects the effect of the thioether bond on the pKa of Y157 and this residue's role in catalysis. At higher pH values, kcat is also higher for the cross-linked form, extending the pH range of activity. We therefore propose that the cross-link also increases activity by controlling deleterious interactions involving the thiol/ate of C93.
- Department of Chemistry and MacDiarmid Institute for Advanced Materials and Nanotechnology and ‡Department of Biochemistry, University of Otago , P.O. Box 56, Dunedin 9054, New Zealand.
Organizational Affiliation: