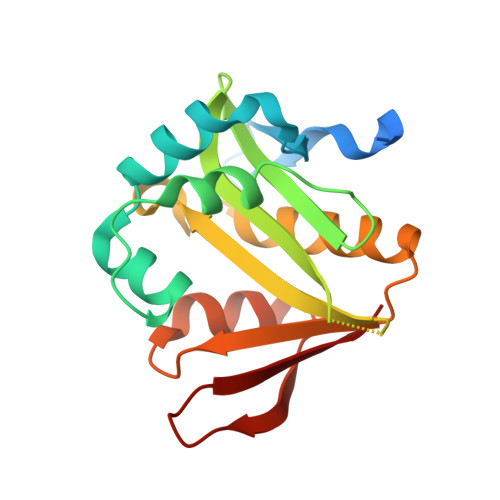
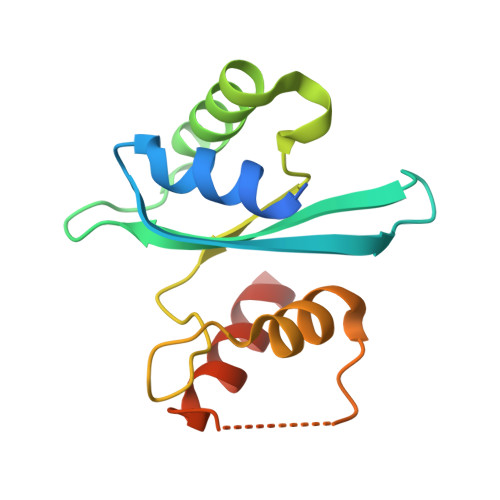
Insights into the specificity of lysine acetyltransferases.
Tucker, A.C., Taylor, K.C., Rank, K.C., Rayment, I., Escalante-Semerena, J.C.(2014) J Biological Chem 289: 36249-36262
- PubMed: 25381442 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.613901
- Primary Citation Related Structures:
4NXY, 4U5Y - PubMed Abstract:
Reversible lysine acetylation by protein acetyltransferases is a conserved regulatory mechanism that controls diverse cellular pathways. Gcn5-related N-acetyltransferases (GNATs), named after their founding member, are found in all domains of life. GNATs are known for their role as histone acetyltransferases, but non-histone bacterial protein acetytransferases have been identified. Only structures of GNAT complexes with short histone peptide substrates are available in databases. Given the biological importance of this modification and the abundance of lysine in polypeptides, how specificity is attained for larger protein substrates is central to understanding acetyl-lysine-regulated networks. Here we report the structure of a GNAT in complex with a globular protein substrate solved to 1.9 Å. GNAT binds the protein substrate with extensive surface interactions distinct from those reported for GNAT-peptide complexes. Our data reveal determinants needed for the recognition of a protein substrate and provide insight into the specificity of GNATs.
- From the Department of Microbiology, University of Georgia, Athens, Georgia 30602 and.
Organizational Affiliation: