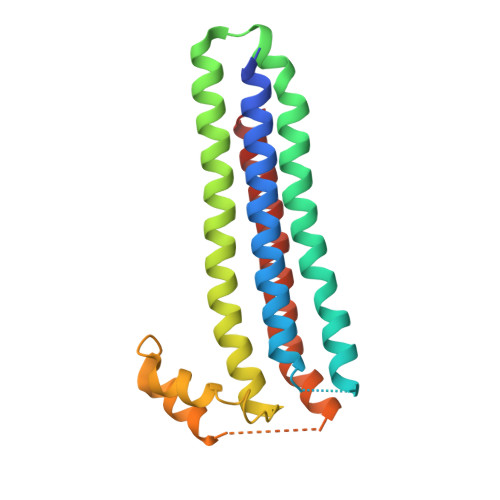
Structure of the Essential Plasmodium Host Cell Traversal Protein SPECT1.
Hamaoka, B.Y., Ghosh, P.(2014) PLoS One 9: e114685-e114685
- PubMed: 25479287 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0114685
- Primary Citation Related Structures:
4U5A - PubMed Abstract:
Host cell traversal by Plasmodium, the protozoan cause of malaria, is an essential part of this parasite's virulence. In this process, the parasite enters a host cell through a parasite-induced pore, traverses the host cell, and then exits the host cell. Two P. berghei proteins, SPECT1 and SPECT2, are required for host cell traversal by the sporozoite form of the parasite. In the absence of either, no pore formation is observed. While SPECT2 has sequence homology to pore-forming proteins, SPECT1 has no homology to proteins of known structure or function. Here we present the 2.75 Å resolution structure of a slightly truncated version of P. berghei SPECT1. The structure reveals that the protein forms a four-helix bundle, with the rare feature of having all of these helices in parallel or antiparallel alignment. Also notable is the presence of a large, conserved, hydrophobic internal cavity in the protein, which may constitute a ligand-binding site or be indicative of partial instability in SPECT1, or both. The structure of SPECT1 will make possible targeted mutagenesis experiments aimed at understanding its mechanism of action in host cell traversal.
- Department of Chemistry & Biochemistry, University of California San Diego, La Jolla, California, United States of America.
Organizational Affiliation: