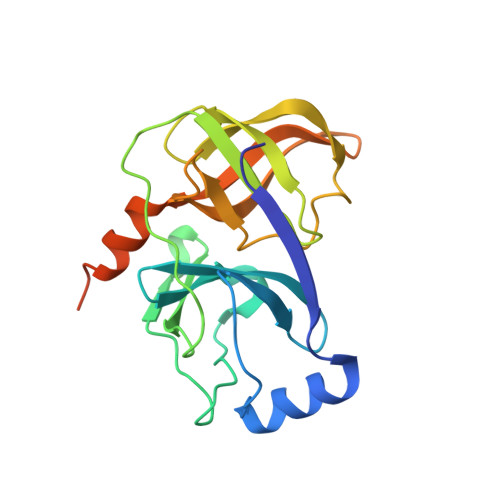
Discovery and structural diversity of the hepatitis C virus NS3/4A serine protease inhibitor series leading to clinical candidate IDX320.
Parsy, C.C., Alexandre, F.R., Bidau, V., Bonnaterre, F., Brandt, G., Caillet, C., Cappelle, S., Chaves, D., Convard, T., Derock, M., Gloux, D., Griffon, Y., Lallos, L.B., Leroy, F., Liuzzi, M., Loi, A.G., Moulat, L., Chiara, M., Rahali, H., Roques, V., Rosinovsky, E., Savin, S., Seifer, M., Standring, D., Surleraux, D.(2015) Bioorg Med Chem Lett 25: 5427-5436
- PubMed: 26410074 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2015.09.009
- Primary Citation Related Structures:
4U01 - PubMed Abstract:
Exploration of the P2 region by mimicking the proline motif found in BILN2061 resulted in the discovery of two series of potent HCV NS3/4A protease inhibitors. X-ray crystal structure of the ligand in contact with the NS3/4A protein and modulation of the quinoline heterocyclic region by structure based design and modeling allowed for the optimization of enzyme potency and cellular activity. This research led to the selection of clinical candidate IDX320 having good genotype coverage and pharmacokinetic properties in various species.
- IDENIX, a wholly-owned subsidiary of Merck & Co, 1682 rue de la Valsière, Cap Gamma, BP 50001, 34189 Montpellier Cedex 4, France. Electronic address: christophe.parsy@merck.com.
Organizational Affiliation: