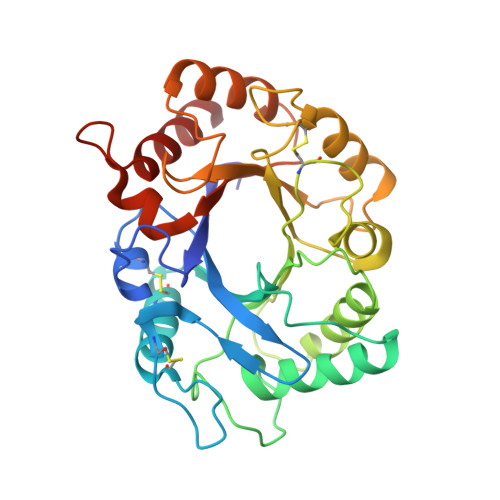
Screening-based discovery of Aspergillus fumigatus plant-type chitinase inhibitors
Lockhart, D.E., Schuettelkopf, A.W., Blair, D.E., van Aalten, D.M.F.(2014) FEBS Lett 588: 3282
- PubMed: 25063338
- DOI: https://doi.org/10.1016/j.febslet.2014.07.015
- Primary Citation of Related Structures:
4TX6, 4TXE - PubMed Abstract:
A limited therapeutic arsenal against increasing clinical disease due to Aspergillus spp. necessitates urgent characterisation of new antifungal targets. Here we describe the discovery of novel, low micromolar chemical inhibitors of Aspergillus fumigatus family 18 plant-type chitinase A1 (AfChiA1) by high-throughput screening (HTS). Analysis of the binding mode by X-ray crystallography confirmed competitive inhibition and kinetic studies revealed two compounds with selectivity towards fungal plant-type chitinases. These inhibitors provide new chemical tools to probe the effects of chitinase inhibition on A. fumigatus growth and virulence, presenting attractive starting points for the development of further potent drug-like molecules.
- Division of Molecular Microbiology, College of Life Sciences, University of Dundee, Dundee DD1 5EH, United Kingdom.
Organizational Affiliation: