2-Benzamido-N-(1H-benzo[d]imidazol-2-yl)thiazole-4-carboxamide derivatives as potent inhibitors of CK1d/e.
Bischof, J., Leban, J., Zaja, M., Grothey, A., Radunsky, B., Othersen, O., Strobl, S., Vitt, D., Knippschild, U.(2012) Amino Acids 43: 1577-1591
- PubMed: 22331384
- DOI: https://doi.org/10.1007/s00726-012-1234-x
- Primary Citation of Related Structures:
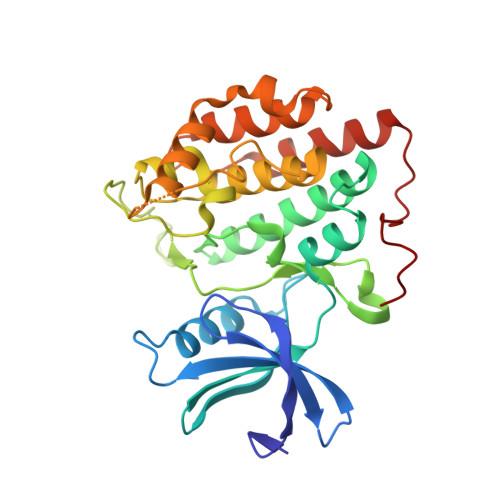
4TWC - PubMed Abstract:
In this study we identified two heterocyclic compounds (5 and 6) as potent and specific inhibitors of CK1δ (IC(50) = 0.040 and 0.042 μM, respectively). Whereas compound 5 exhibited fivefold higher affinity towards CK1δ than to CK1ε (IC(50) CK1ε = 0.199 μM), compound 6 also inhibited CK1ε (IC(50) = 0.0326 μM) in the same range as CK1δ. Selected compound 5 was screened over 442 kinases identifying 5 as a highly potent and selective inhibitor of CK1δ. X-ray analysis of 5 bound to CK1δ demonstrated its binding mode. In addition, characterization of 5 and 6 in a cell biological approach revealed the ability of both compounds to inhibit proliferation of tumor cell lines in a dose and cell line specific manner. In summary, our optimizations lead to the development of new highly selective CK1δ and ε specific inhibitors with biological activity.
- Department of General, Visceral and Transplantation Surgery, Surgery Centre, University of Ulm, Steinhövel Str. 9, 89075, Ulm, Germany.
Organizational Affiliation: