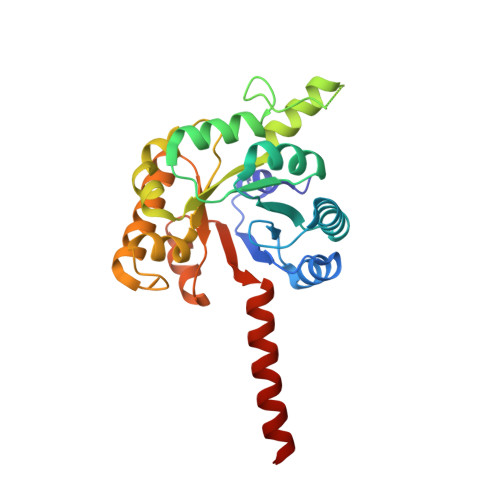
SbnG, a Citrate Synthase in Staphylococcus aureus: A NEW FOLD ON AN OLD ENZYME.
Kobylarz, M.J., Grigg, J.C., Sheldon, J.R., Heinrichs, D.E., Murphy, M.E.(2014) J Biological Chem 289: 33797-33807
- PubMed: 25336653
- DOI: https://doi.org/10.1074/jbc.M114.603175
- Primary Citation Related Structures:
4TV5, 4TV6 - PubMed Abstract:
In response to iron deprivation, Staphylococcus aureus produces staphyloferrin B, a citrate-containing siderophore that delivers iron back to the cell. This bacterium also possesses a second citrate synthase, SbnG, that is necessary for supplying citrate to the staphyloferrin B biosynthetic pathway. We present the structure of SbnG bound to the inhibitor calcium and an active site variant in complex with oxaloacetate. The overall fold of SbnG is structurally distinct from TCA cycle citrate synthases yet similar to metal-dependent class II aldolases. Phylogenetic analyses revealed that SbnG forms a separate clade with homologs from other siderophore biosynthetic gene clusters and is representative of a metal-independent subgroup in the phosphoenolpyruvate/pyruvate domain superfamily. A structural superposition of the SbnG active site to TCA cycle citrate synthases and site-directed mutagenesis suggests a case for convergent evolution toward a conserved catalytic mechanism for citrate production.
- From the Department of Microbiology and Immunology, Life Sciences Institute, University of British Columbia, Vancouver, British Columbia V6T 1Z3, Canada and.
Organizational Affiliation: