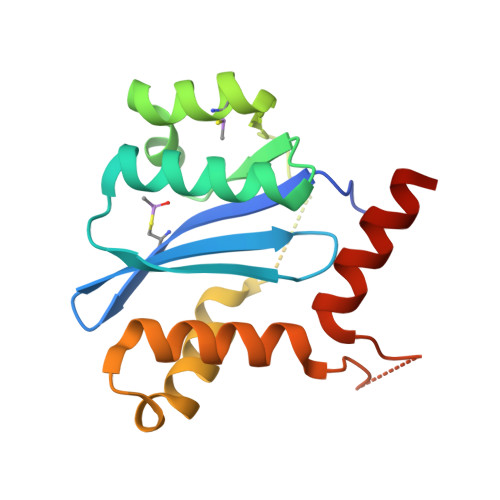
The mechanism of H171T resistance reveals the importance of N -protonated His171 for the binding of allosteric inhibitor BI-D to HIV-1 integrase.
Slaughter, A., Jurado, K.A., Deng, N., Feng, L., Kessl, J.J., Shkriabai, N., Larue, R.C., Fadel, H.J., Patel, P.A., Jena, N., Fuchs, J.R., Poeschla, E., Levy, R.M., Engelman, A., Kvaratskhelia, M.(2014) Retrovirology 11: 100-100
- PubMed: 25421939
- DOI: https://doi.org/10.1186/s12977-014-0100-1
- Primary Citation of Related Structures:
4TSX - PubMed Abstract:
Allosteric HIV-1 integrase (IN) inhibitors (ALLINIs) are an important new class of anti-HIV-1 agents. ALLINIs bind at the IN catalytic core domain (CCD) dimer interface occupying the principal binding pocket of its cellular cofactor LEDGF/p75. Consequently, ALLINIs inhibit HIV-1 IN interaction with LEDGF/p75 as well as promote aberrant IN multimerization. Selection of viral strains emerging under the inhibitor pressure has revealed mutations at the IN dimer interface near the inhibitor binding site. We have investigated the effects of one of the most prevalent substitutions, H171T IN, selected under increasing pressure of ALLINI BI-D. Virus containing the H171T IN substitution exhibited an ~68-fold resistance to BI-D treatment in infected cells. These results correlated with ~84-fold reduced affinity for BI-D binding to recombinant H171T IN CCD protein compared to its wild type (WT) counterpart. However, the H171T IN substitution only modestly affected IN-LEDGF/p75 binding and allowed HIV-1 containing this substitution to replicate at near WT levels. The x-ray crystal structures of BI-D binding to WT and H171T IN CCD dimers coupled with binding free energy calculations revealed the importance of the Nδ- protonated imidazole group of His171 for hydrogen bonding to the BI-D tert-butoxy ether oxygen and establishing electrostatic interactions with the inhibitor carboxylic acid, whereas these interactions were compromised upon substitution to Thr171. Our findings reveal a distinct mechanism of resistance for the H171T IN mutation to ALLINI BI-D and indicate a previously undescribed role of the His171 side chain for binding the inhibitor.
- Center for Retrovirus Research and Comprehensive Cancer Center, College of Pharmacy, The Ohio State University, 496 W. 12th Ave, 508 Riffe Building, Columbus, OH, 43210, USA. slaughter.89@osu.edu.
Organizational Affiliation: