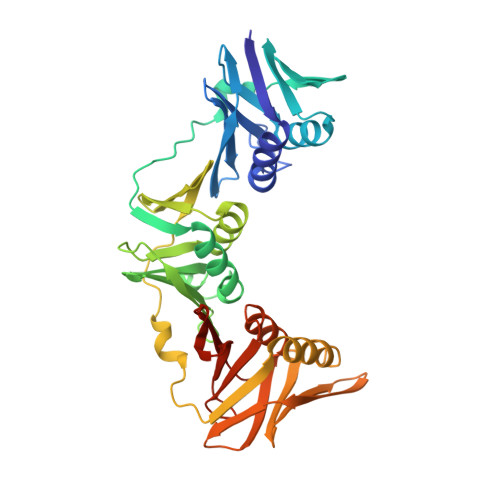
Crystal structure of the DNA polymerase III beta subunit ( beta-clamp) from the extremophile Deinococcus radiodurans.
Niiranen, L., Lian, K., Johnson, K.A., Moe, E.(2015) BMC Struct Biol 15: 5-5
- PubMed: 25886944
- DOI: https://doi.org/10.1186/s12900-015-0032-6
- Primary Citation Related Structures:
4TRT - PubMed Abstract:
Deinococcus radiodurans is an extremely radiation and desiccation resistant bacterium which can tolerate radiation doses up to 5,000 Grays without losing viability. We are studying the role of DNA repair and replication proteins for this unusual phenotype by a structural biology approach. The DNA polymerase III β subunit (β-clamp) acts as a sliding clamp on DNA, promoting the binding and processivity of many DNA-acting proteins, and here we report the crystal structure of D. radiodurans β-clamp (Drβ-clamp) at 2.0 Å resolution. The sequence verification process revealed that at the time of the study the gene encoding Drβ-clamp was wrongly annotated in the genome database, encoding a protein of 393 instead of 362 amino acids. The short protein was successfully expressed, purified and used for crystallisation purposes in complex with Cy5-labeled DNA. The structure, which was obtained from blue crystals, shows a typical ring-shaped bacterial β-clamp formed of two monomers, each with three domains of identical topology, but with no visible DNA in electron density. A visualisation of the electrostatic surface potential reveals a highly negatively charged outer surface while the inner surface and the dimer forming interface have a more even charge distribution. The structure of Drβ-clamp was determined to 2.0 Å resolution and shows an evenly distributed electrostatic surface charge on the DNA interacting side. We hypothesise that this charge distribution may facilitate efficient movement on encircled DNA and help ensure efficient DNA metabolism in D. radiodurans upon exposure to high doses of ionizing irradiation or desiccation.
- The Norwegian Structural Biology Center (NorStruct), Department of Chemistry, UIT - the Arctic University of Norway, N-9037, Tromsø, Norway. laila.niiranen@live.fi.
Organizational Affiliation: