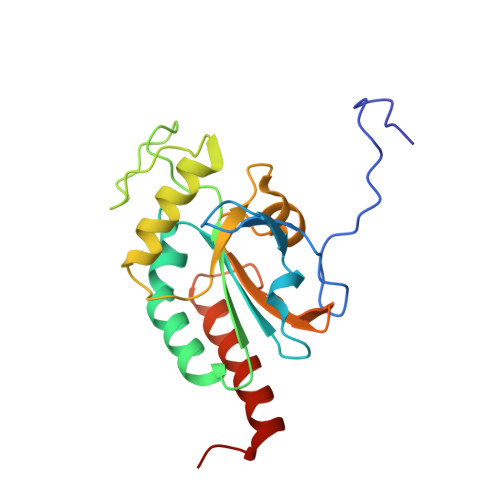
Crystal structure of mammalian selenocysteine-dependent iodothyronine deiodinase suggests a peroxiredoxin-like catalytic mechanism.
Schweizer, U., Schlicker, C., Braun, D., Kohrle, J., Steegborn, C.(2014) Proc Natl Acad Sci U S A 111: 10526
- PubMed: 25002520 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1323873111
- Primary Citation Related Structures:
4TR3, 4TR4 - PubMed Abstract:
Local levels of active thyroid hormone (3,3',5-triiodothyronine) are controlled by the action of activating and inactivating iodothyronine deiodinase enzymes. Deiodinases are selenocysteine-dependent membrane proteins catalyzing the reductive elimination of iodide from iodothyronines through a poorly understood mechanism. We solved the crystal structure of the catalytic domain of mouse deiodinase 3 (Dio3), which reveals a close structural similarity to atypical 2-Cys peroxiredoxin(s) (Prx). The structure suggests a route for proton transfer to the substrate during deiodination and a Prx-related mechanism for subsequent recycling of the transiently oxidized enzyme. The proposed mechanism is supported by biochemical experiments and is consistent with the effects of mutations of conserved amino acids on Dio3 activity. Thioredoxin and glutaredoxin reduce the oxidized Dio3 at physiological concentrations, and dimerization appears to activate the enzyme by displacing an autoinhibitory loop from the iodothyronine binding site. Deiodinases apparently evolved from the ubiquitous Prx scaffold, and their structure and catalytic mechanism reconcile a plethora of partly conflicting data reported for these enzymes.
- Institut für Biochemie und Molekularbiologie, Rheinische Friedrich Wilhelms-Universität Bonn, 53115 Bonn, Germany; uschweiz@uni-bonn.de clemens.steegborn@uni-bayreuth.de.
Organizational Affiliation: