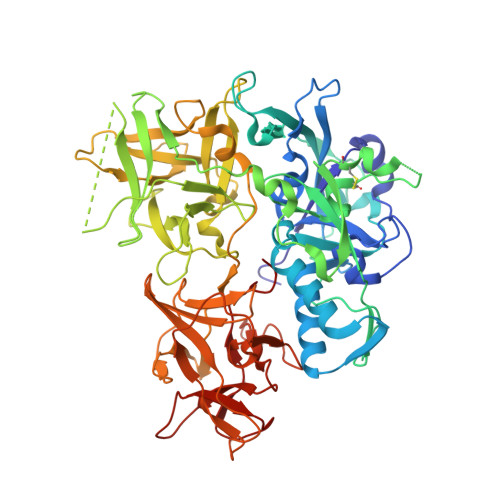
Structure of CARDS toxin, a unique ADP-ribosylating and vacuolating cytotoxin from Mycoplasma pneumoniae.
Becker, A., Kannan, T.R., Taylor, A.B., Pakhomova, O.N., Zhang, Y., Somarajan, S.R., Galaleldeen, A., Holloway, S.P., Baseman, J.B., Hart, P.J.(2015) Proc Natl Acad Sci U S A 112: 5165-5170
- PubMed: 25848012 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1420308112
- Primary Citation Related Structures:
4TLV, 4TLW - PubMed Abstract:
Mycoplasma pneumoniae (Mp) infections cause tracheobronchitis and "walking" pneumonia, and are linked to asthma and other reactive airway diseases. As part of the infectious process, the bacterium expresses a 591-aa virulence factor with both mono-ADP ribosyltransferase (mART) and vacuolating activities known as Community-Acquired Respiratory Distress Syndrome Toxin (CARDS TX). CARDS TX binds to human surfactant protein A and annexin A2 on airway epithelial cells and is internalized, leading to a range of pathogenetic events. Here we present the structure of CARDS TX, a triangular molecule in which N-terminal mART and C-terminal tandem β-trefoil domains associate to form an overall architecture distinct from other well-recognized ADP-ribosylating bacterial toxins. We demonstrate that CARDS TX binds phosphatidylcholine and sphingomyelin specifically over other membrane lipids, and that cell surface binding and internalization activities are housed within the C-terminal β-trefoil domain. The results enhance our understanding of Mp pathogenicity and suggest a novel avenue for the development of therapies to treat Mp-associated asthma and other acute and chronic airway diseases.
- Department of Biochemistry.
Organizational Affiliation: