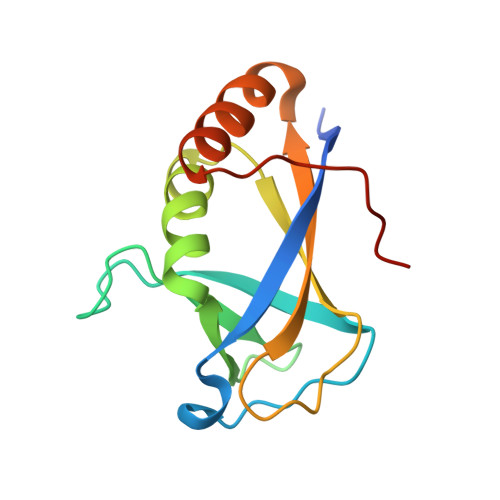
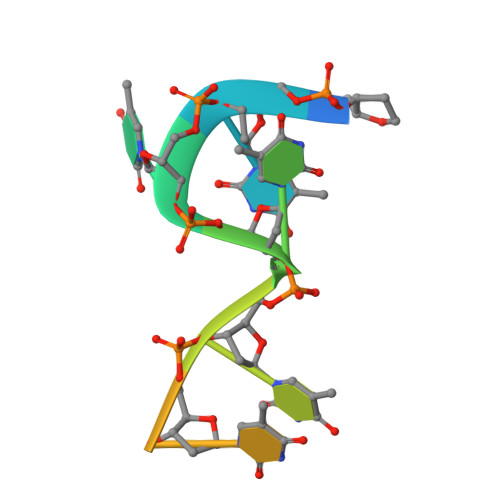
HLTF's Ancient HIRAN Domain Binds 3' DNA Ends to Drive Replication Fork Reversal.
Kile, A.C., Chavez, D.A., Bacal, J., Eldirany, S., Korzhnev, D.M., Bezsonova, I., Eichman, B.F., Cimprich, K.A.(2015) Mol Cell 58: 1090-1100
- PubMed: 26051180 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2015.05.013
- Primary Citation Related Structures:
2MZN, 4S0N - PubMed Abstract:
Stalled replication forks are a critical problem for the cell because they can lead to complex genome rearrangements that underlie cell death and disease. Processes such as DNA damage tolerance and replication fork reversal protect stalled forks from these events. A central mediator of these DNA damage responses in humans is the Rad5-related DNA translocase, HLTF. Here, we present biochemical and structural evidence that the HIRAN domain, an ancient and conserved domain found in HLTF and other DNA processing proteins, is a modified oligonucleotide/oligosaccharide (OB) fold that binds to 3' ssDNA ends. We demonstrate that the HIRAN domain promotes HLTF-dependent fork reversal in vitro through its interaction with 3' ssDNA ends found at forks. Finally, we show that HLTF restrains replication fork progression in cells in a HIRAN-dependent manner. These findings establish a mechanism of HLTF-mediated fork reversal and provide insight into the requirement for distinct fork remodeling activities in the cell.
- Department of Chemical and Systems Biology, Stanford University School of Medicine, Stanford, CA 94305, USA.
Organizational Affiliation: