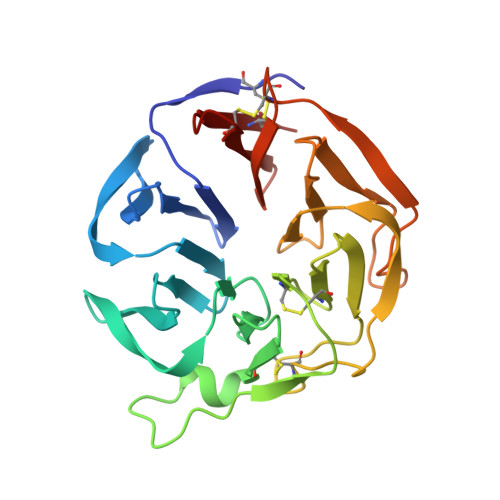
Three-dimensional structure and ligand-binding site of carp fishelectin (FEL).
Capaldi, S., Faggion, B., Carrizo, M.E., Destefanis, L., Gonzalez, M.C., Perduca, M., Bovi, M., Galliano, M., Monaco, H.L.(2015) Acta Crystallogr D Biol Crystallogr 71: 1123-1135
- PubMed: 25945578 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004715004174
- Primary Citation Related Structures:
4RUQ, 4RUS - PubMed Abstract:
Carp FEL (fishelectin or fish-egg lectin) is a 238-amino-acid lectin that can be purified from fish eggs by exploiting its selective binding to Sepharose followed by elution with N-acetylglucosamine. Its amino-acid sequence and other biochemical properties have previously been reported. The glycoprotein has four disulfide bridges and the structure of the oligosaccharides linked to Asn27 has been described. Here, the three-dimensional structures of apo carp FEL (cFEL) and of its complex with N-acetylglucosamine determined by X-ray crystallography at resolutions of 1.35 and 1.70 Å, respectively, are reported. The molecule folds as a six-bladed β-propeller and internal short consensus amino-acid sequences have been identified in all of the blades. A calcium atom binds at the bottom of the funnel-shaped tunnel located in the centre of the propeller. Two ligand-binding sites, α and β, are present in each of the two protomers in the dimer. The first site, α, is closer to the N-terminus of the chain and is located in the crevice between the second and the third blades, while the second site, β, is located between the fourth and the fifth blades. The amino acids that participate in the contacts have been identified, as well as the conserved water molecules in all of the sites. Both sites can bind the two anomers, α and β, of N-acetylglucosamine, as is clearly recognizable in the electron-density maps. The lectin presents sequence homology to members of the tachylectin family, which are known to have a function in the innate immune system of arthropods, and homologous genes are present in the genomes of other fish and amphibians. This structure is the first of a protein of this group and, given the degree of homology with other members of the family, it is expected that it will be useful to experimentally determine other crystal structures using the coordinates of cFEL as a search probe in molecular replacement.
- Biocrystallography Laboratory, Department of Biotechnology, University of Verona, Ca Vignal 1, Strada Le Grazie 15, 37134 Verona, Italy.
Organizational Affiliation: