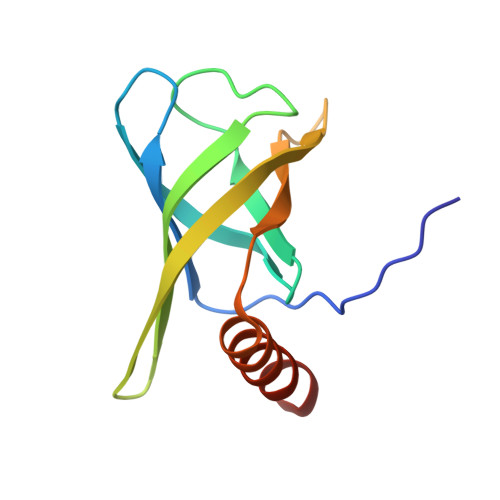
Dimeric c-di-GMP Is Required for Post-translational Regulation of Alginate Production in Pseudomonas aeruginosa.
Whitney, J.C., Whitfield, G.B., Marmont, L.S., Yip, P., Neculai, A.M., Lobsanov, Y.D., Robinson, H., Ohman, D.E., Howell, P.L.(2015) J Biological Chem 290: 12451-12462
- PubMed: 25817996 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.645051
- Primary Citation Related Structures:
4RT0, 4RT1 - PubMed Abstract:
Pseudomonas aeruginosa is an opportunistic human pathogen that secretes the exopolysaccharide alginate during infection of the respiratory tract of individuals afflicted with cystic fibrosis and chronic obstructive pulmonary disease. Among the proteins required for alginate production, Alg44 has been identified as an inner membrane protein whose bis-(3',5')-cyclic dimeric guanosine monophosphate (c-di-GMP) binding activity post-translationally regulates alginate secretion. In this study, we report the 1.8 Å crystal structure of the cytoplasmic region of Alg44 in complex with dimeric self-intercalated c-di-GMP and characterize its dinucleotide-binding site using mutational analysis. The structure shows that the c-di-GMP binding region of Alg44 adopts a PilZ domain fold with a dimerization mode not previously observed for this family of proteins. Calorimetric binding analysis of residues in the c-di-GMP binding site demonstrate that mutation of Arg-17 and Arg-95 alters the binding stoichiometry between c-di-GMP and Alg44 from 2:1 to 1:1. Introduction of these mutant alleles on the P. aeruginosa chromosome show that the residues required for binding of dimeric c-di-GMP in vitro are also required for efficient alginate production in vivo. These results suggest that the dimeric form of c-di-GMP represents the biologically active signaling molecule needed for the secretion of an important virulence factor produced by P. aeruginosa.
- From the Program in Molecular Structure and Function, Hospital for Sick Children, Toronto, Ontario M5G 0A4, Canada, the Department of Biochemistry, University of Toronto, Toronto, Ontario M5S 1A8, Canada.
Organizational Affiliation: