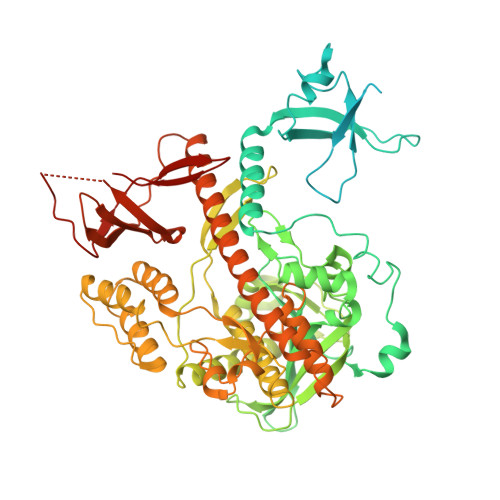
Structural analysis of Dis3l2, an exosome-independent exonuclease from Schizosaccharomyces pombe.
Lv, H., Zhu, Y., Qiu, Y., Niu, L., Teng, M., Li, X.(2015) Acta Crystallogr D Biol Crystallogr 71: 1284-1294
- PubMed: 26057668 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004715005805
- Primary Citation Related Structures:
4RO1 - PubMed Abstract:
After deadenylation and decapping, cytoplasmic mRNA can be digested in two opposite directions: in the 5'-3' direction by Xrn1 or in the 3'-5' direction by the exosome complex. Recently, a novel 3'-5' RNA-decay pathway involving Dis3l2 has been described that differs from degradation by Xrn1 and the exosome. The product of the Schizosaccharomyces pombe gene SPAC2C4.07c was identified as a homologue of human Dis3l2. In this work, the 2.8 Å resolution X-ray crystal structure of S. pombe Dis3l2 (SpDis3l2) is reported, the conformation of which is obviously different from that in the homologous mouse Dis3l2-RNA complex. Fluorescence polarization assay experiments showed that RNB and S1 are the primary RNA-binding domains and that the CSDs (CSD1 and CSD2) play an indispensable role in the RNA-binding process of SpDis3l2. Taking the structure comparison and mutagenic experiments together, it can be inferred that the RNA-recognition pattern of SpDis3l2 resembles that of its mouse homologue rather than that of the Escherichia coli RNase II-RNA complex. Furthermore, a drastic conformation change could occur following the binding of the RNA substrate to SpDis3l2.
- Hefei National Laboratory for Physical Sciences at Microscale and School of Life Science, University of Science and Technology of China, Hefei, Anhui 230026, People's Republic of China.
Organizational Affiliation: