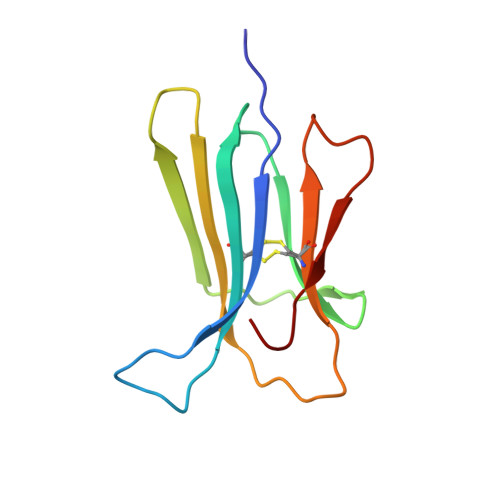
Decoding the Structural Bases of D76N 2-Microglobulin High Amyloidogenicity through Crystallography and Asn-Scan Mutagenesis.
de Rosa, M., Barbiroli, A., Giorgetti, S., Mangione, P.P., Bolognesi, M., Ricagno, S.(2015) PLoS One 10: e0144061-e0144061
- PubMed: 26625273 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0144061
- Primary Citation Related Structures:
4RMQ, 4RMR, 4RMS, 4RMT - PubMed Abstract:
D76N is the first natural variant of human β-2 microglobulin (β2m) so far identified. Contrary to the wt protein, this mutant readily forms amyloid fibres in physiological conditions, leading to a systemic and severe amyloidosis. Although the Asp76Asn mutant has been extensively characterized, the molecular bases of its instability and aggregation propensity remain elusive. In this work all Asp residues of human β2m were individually substituted to Asn; D-to-N mutants (D34N, D38N, D53N, D59N, D96N and D98N) were characterised in terms of thermodynamic stability and aggregation propensity. Moreover, crystal structures of the D38N, D53N, D59N and D98N variants were solved at high-resolution (1.24-1.70 Å). Despite showing some significant variations in their thermal stabilities, none showed the dramatic drop in melting temperature (relative to the wt protein) as observed for the pathogenic mutant. Consistently, none of the variants here described displayed any increase in aggregation propensity under the experimental conditions tested. The crystal structures confirmed that D-to-N mutations are generally well tolerated, and lead only to minor reorganization of the side chains in close proximity of the mutated residue. D38N is the only exception, where backbone readjustments and a redistribution of the surface electrostatic charges are observed. Overall, our results suggest that neither removing negative charges at sites 34, 38, 53, 59, 96 and 98, nor the difference in β2m pI, are the cause of the aggressive phenotype observed in D76N. We propose that the dramatic effects of the D76N natural mutation must be linked to effects related to the crucial location of this residue within the β2m fold.
- Dipartimento di Bioscienze, Università di Milano, Via Celoria 26, 20133, Milano, Italy.
Organizational Affiliation: