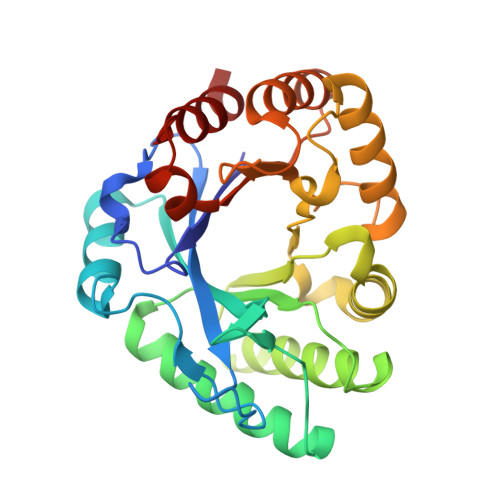
A class III chitinase without disulfide bonds from the fern, Pteris ryukyuensis: crystal structure and ligand-binding studies.
Kitaoku, Y., Umemoto, N., Ohnuma, T., Numata, T., Taira, T., Sakuda, S., Fukamizo, T.(2015) Planta 242: 895-907
- PubMed: 25998529 Search on PubMed
- DOI: https://doi.org/10.1007/s00425-015-2330-4
- Primary Citation Related Structures:
4RL3 - PubMed Abstract:
We first solved the crystal structure of class III catalytic domain of a chitinase from fern (PrChiA-cat), and found a structural difference between PrChiA-cat and hevamine. PrChiA-cat was found to have reduced affinities to chitin oligosaccharides and allosamidin. Plant class III chitinases are subdivided into enzymes with three disulfide bonds and those without disulfide bonds. We here referred to the former enzymes as class IIIa chitinases and the latter as class IIIb chitinases. In this study, we solved the crystal structure of the class IIIb catalytic domain of a chitinase from the fern Pteris ryukyuensis (PrChiA-cat), and compared it with that of hevamine, a class IIIa chitinase from Hevea brasiliensis. PrChiA-cat was found to adopt an (α/β)8 fold typical of GH18 chitinases in a similar manner to that of hevamine. However, PrChiA-cat also had two large loops that extruded from the catalytic site, and the corresponding loops in hevamine were markedly smaller than those of PrChiA-cat. An HPLC analysis of the enzymatic products revealed that the mode of action of PrChiA-cat toward chitin oligosaccharides, (GlcNAc) n (n = 4-6), differed from those of hevamine and the other class IIIa chitinases. The binding affinities of (GlcNAc)3 and (GlcNAc)4 toward the inactive mutant of PrChiA-cat were determined by isothermal titration calorimetry, and were markedly lower than those toward other members of the GH18 family. The affinity and the inhibitory activity of allosamidin toward PrChiA-cat were also lower than those toward the GH18 chitinases investigated to date. Several hydrogen bonds found in the crystal structure of hevamine-allosamidin complex were missing in the modeled structure of PrChiA-cat-allosamidin complex. The structural findings for PrChiA-cat successfully interpreted the functional data presented.
- Department of Advanced Bioscience, Kinki University, 3327-204 Nakamachi, Nara, 631-8505, Japan.
Organizational Affiliation: