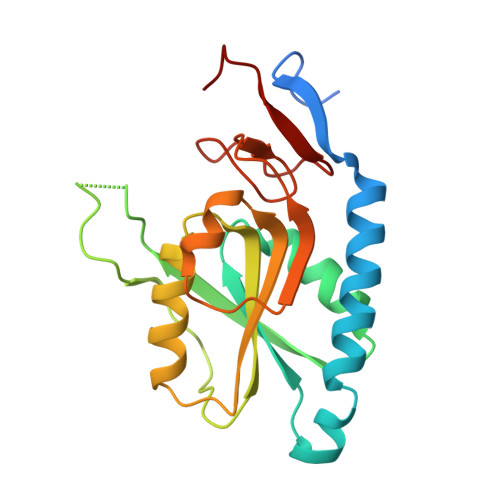
First Crystal Structures of Mycobacterium tuberculosis 6-Oxopurine Phosphoribosyltransferase: Complexes with GMP and Pyrophosphate and with Acyclic Nucleoside Phosphonates Whose Prodrugs Have Antituberculosis Activity.
Eng, W.S., Hockova, D., Spacek, P., Janeba, Z., West, N.P., Woods, K., Naesens, L.M., Keough, D.T., Guddat, L.W.(2015) J Med Chem 58: 4822-4838
- PubMed: 25915781 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.5b00611
- Primary Citation Related Structures:
4RHT, 4RHU, 4RHX, 4RHY - PubMed Abstract:
Human tuberculosis is a chronic infectious disease affecting millions of lives. Because of emerging resistance to current medications, new therapeutic drugs are needed. One potential new target is hypoxanthine-guanine phosphoribosyltransferase (MtHGPRT), a key enzyme of the purine salvage pathway. Here, newly synthesized acyclic nucleoside phosphonates (ANPs) have been shown to be competitive inhibitors of MtHGPRT with Ki values as low as 0.69 μM. Prodrugs of these compounds arrest the growth of a virulent strain of M. tuberculosis with MIC50 values as low as 4.5 μM and possess low cytotoxicity in mammalian cells (CC50 values as high as >300 μM). In addition, the first crystal structures of MtHGPRT (2.03-2.76 Å resolution) have been determined, three of these in complex with novel ANPs and one with GMP and pyrophosphate. These data provide a solid foundation for the further development of ANPs as selective inhibitors of MtHGPRT and as antituberculosis agents.
- †The School of Chemistry and Molecular Biosciences, The University of Queensland, Brisbane, 4072 QLD Australia.
Organizational Affiliation: