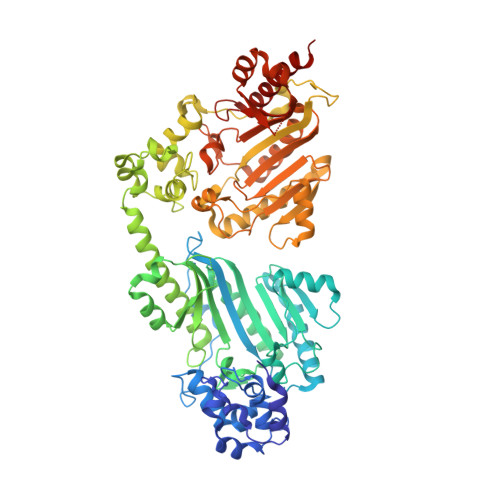
Structure of a double-domain phosphagen kinase reveals an asymmetric arrangement of the tandem domains.
Wang, Z., Qiao, Z., Ye, S., Zhang, R.(2015) Acta Crystallogr D Biol Crystallogr 71: 779-789
- PubMed: 25849389 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004715001169
- Primary Citation Related Structures:
4RF6, 4RF7, 4RF8, 4RF9 - PubMed Abstract:
Tandem duplications and fusions of single genes have led to magnificent expansions in the divergence of protein structures and functions over evolutionary timescales. One of the possible results is polydomain enzymes with interdomain cooperativities, few examples of which have been structurally characterized at the full-length level to explore their innate synergistic mechanisms. This work reports the crystal structures of a double-domain phosphagen kinase in both apo and ligand-bound states, revealing a novel asymmetric L-shaped arrangement of the two domains. Unexpectedly, the interdomain connections are not based on a flexible hinge linker but on a rigid secondary-structure element: a long α-helix that tethers the tandem domains in relatively fixed positions. Besides the connective helix, the two domains also contact each other directly and form an interdomain interface in which hydrogen bonds and hydrophobic interactions further stabilize the L-shaped domain arrangement. Molecular-dynamics simulations show that the interface is generally stable, suggesting that the asymmetric domain arrangement crystallographically observed in the present study is not a conformational state simply restrained by crystal-packing forces. It is possible that the asymmetrically arranged tandem domains could provide a structural basis for further studies of the interdomain synergy.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, People's Republic of China.
Organizational Affiliation: