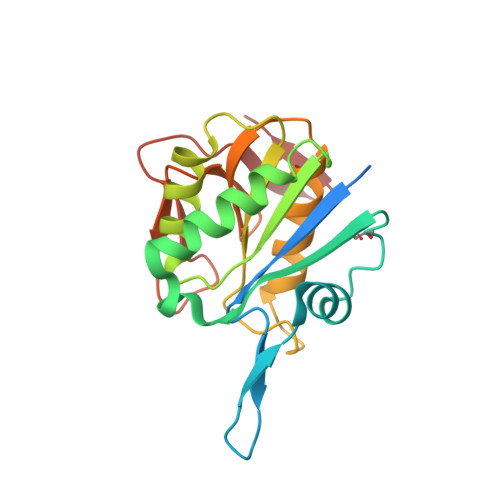
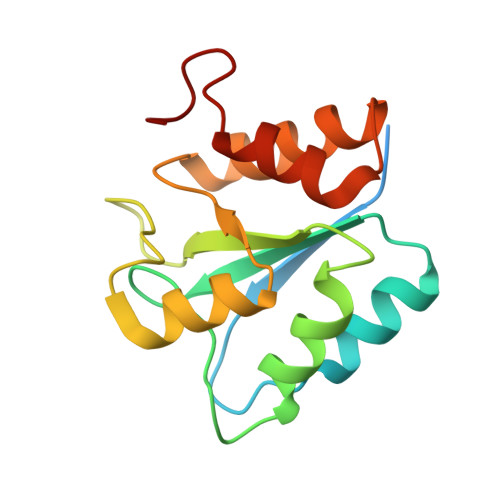
Structural and functional insights into an archaeal L-asparaginase obtained through the linker-less assembly of constituent domains.
Tomar, R., Sharma, P., Srivastava, A., Bansal, S., Kundu, B.(2014) Acta Crystallogr D Biol Crystallogr 70: 3187-3197
- PubMed: 25478837
- DOI: https://doi.org/10.1107/S1399004714023414
- Primary Citation of Related Structures:
4NJE, 4Q0M, 4RA6, 4RA9 - PubMed Abstract:
Covalent linkers bridging the domains of multidomain proteins are considered to be crucial for assembly and function. In this report, an exception in which the linker of a two-domain dimeric L-asparaginase from Pyrococcus furiosus (PfA) was found to be dispensable is presented. Domains of this enzyme assembled without the linker into a conjoined tetrameric form that exhibited higher activity than the parent enzyme. The global shape and quaternary structure of the conjoined PfA were also similar to the wild-type PfA, as observed by their solution scattering profiles and X-ray crystallographic data. Comparison of the crystal structures of substrate-bound and unbound enzymes revealed an altogether new active-site composition and mechanism of action. Thus, conjoined PfA is presented as a unique enzyme obtained through noncovalent, linker-less assembly of constituent domains that is stable enough to function efficiently at elevated temperatures.
- Kusuma School of Biological Sciences, Indian Institute of Technology Delhi, New Delhi, India.
Organizational Affiliation: