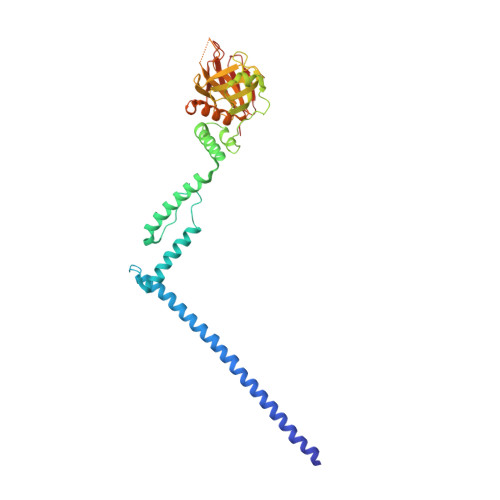
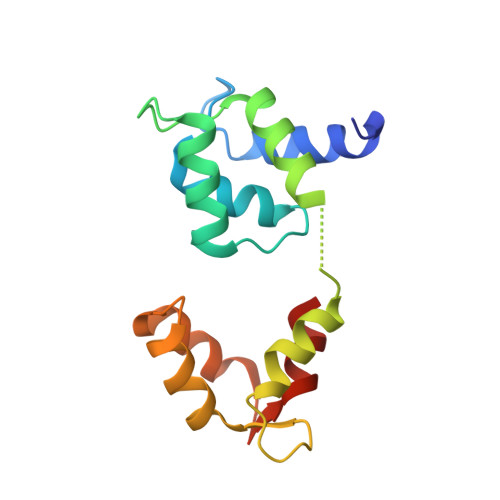
Structure of myosin-1c tail bound to calmodulin provides insights into calcium-mediated conformational coupling.
Lu, Q., Li, J., Ye, F., Zhang, M.(2015) Nat Struct Mol Biol 22: 81-88
- PubMed: 25437912 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2923
- Primary Citation Related Structures:
4R8G - PubMed Abstract:
Class I myosins can sense cellular mechanical forces and function as tension-sensitive anchors or transporters. How mechanical load is transduced from the membrane-binding tail to the force-generating head in myosin-1 is unknown. Here we determined the crystal structure of the entire tail of mouse myosin-1c in complex with apocalmodulin, showing that myosin-1c adopts a stable monomer conformation suited for force transduction. The lever-arm helix and the C-terminal extended PH domain of the motor are coupled by a stable post-IQ domain bound to calmodulin in a highly unusual mode. Ca(2+) binding to calmodulin induces major conformational changes in both IQ motifs and the post-IQ domain and increases flexibility of the myosin-1c tail. Our study provides a structural blueprint for the neck and tail domains of myosin-1 and expands the target binding modes of the master Ca(2+)-signal regulator calmodulin.
- Division of Life Science, Hong Kong University of Science and Technology, Clear Water Bay, Kowloon, Hong Kong, China.
Organizational Affiliation: