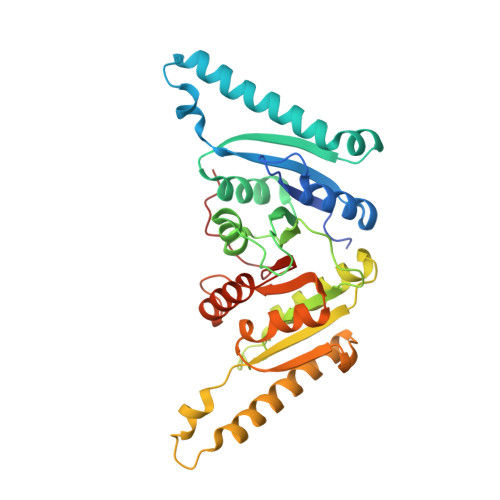
Structural and functional analysis of two universal stress proteins YdaA and YnaF from Salmonella typhimurium: possible roles in microbial stress tolerance.
Bangera, M., Panigrahi, R., Sagurthi, S.R., Savithri, H.S., Murthy, M.R.(2015) J Struct Biol 189: 238-250
- PubMed: 25600413
- DOI: https://doi.org/10.1016/j.jsb.2015.01.003
- Primary Citation of Related Structures:
4R2J, 4R2K, 4R2L, 4R2M - PubMed Abstract:
In many organisms "Universal Stress Proteins" (USPs) are induced in response to a variety of environmental stresses. Here we report the structures of two USPs, YnaF and YdaA from Salmonella typhimurium determined at 1.8Å and 2.4Å resolutions, respectively. YnaF consists of a single USP domain and forms a tetrameric organization stabilized by interactions mediated through chloride ions. YdaA is a larger protein consisting of two tandem USP domains. Two protomers of YdaA associate to form a structure similar to the YnaF tetramer. YdaA showed ATPase activity and an ATP binding motif G-2X-G-9X-G(S/T/N) was found in its C-terminal domain. The residues corresponding to this motif were not conserved in YnaF although YnaF could bind ATP. However, unlike YdaA, YnaF did not hydrolyse ATP in vitro. Disruption of interactions mediated through chloride ions by selected mutations converted YnaF into an ATPase. Residues that might be important for ATP hydrolysis could be identified by comparing the active sites of native and mutant structures. Only the C-terminal domain of YdaA appears to be involved in ATP hydrolysis. The structurally similar N-terminal domain was found to bind a zinc ion near the segment equivalent to the phosphate binding loop of the C-terminal domain. Mass spectrometric analysis showed that YdaA might bind a ligand of approximate molecular weight 800daltons. Structural comparisons suggest that the ligand, probably related to an intermediate in lipid A biosynthesis, might bind at a site close to the zinc ion. Therefore, the N-terminal domain of YdaA binds zinc and might play a role in lipid metabolism. Thus, USPs appear to perform several distinct functions such as ATP hydrolysis, altering membrane properties and chloride sensing.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore 560012, India.
Organizational Affiliation: