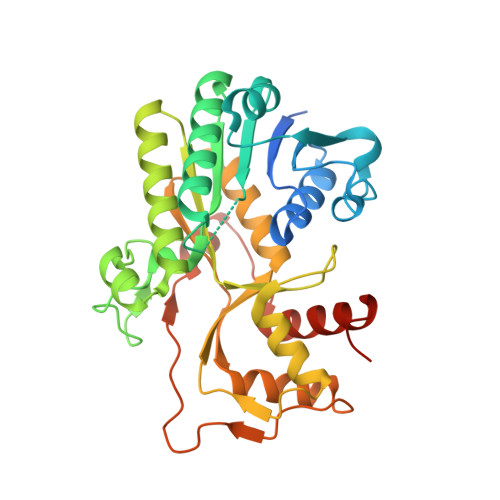
Structural Studies of Cinnamoyl-CoA Reductase and Cinnamyl-Alcohol Dehydrogenase, Key Enzymes of Monolignol Biosynthesis.
Pan, H., Zhou, R., Louie, G.V., Muhlemann, J.K., Bomati, E.K., Bowman, M.E., Dudareva, N., Dixon, R.A., Noel, J.P., Wang, X.(2014) Plant Cell 26: 3709-3727
- PubMed: 25217505
- DOI: https://doi.org/10.1105/tpc.114.127399
- Primary Citation Related Structures:
4QTZ, 4QUK, 4R1S, 4R1T, 4R1U - PubMed Abstract:
The enzymes cinnamoyl-CoA reductase (CCR) and cinnamyl alcohol dehydrogenase (CAD) catalyze the two key reduction reactions in the conversion of cinnamic acid derivatives into monolignol building blocks for lignin polymers in plant cell walls. Here, we describe detailed functional and structural analyses of CCRs from Medicago truncatula and Petunia hybrida and of an atypical CAD (CAD2) from M. truncatula. These enzymes are closely related members of the short-chain dehydrogenase/reductase (SDR) superfamily. Our structural studies support a reaction mechanism involving a canonical SDR catalytic triad in both CCR and CAD2 and an important role for an auxiliary cysteine unique to CCR. Site-directed mutants of CAD2 (Phe226Ala and Tyr136Phe) that enlarge the phenolic binding site result in a 4- to 10-fold increase in activity with sinapaldehyde, which in comparison to the smaller coumaraldehyde and coniferaldehyde substrates is disfavored by wild-type CAD2. This finding demonstrates the potential exploitation of rationally engineered forms of CCR and CAD2 for the targeted modification of monolignol composition in transgenic plants. Thermal denaturation measurements and structural comparisons of various liganded and unliganded forms of CCR and CAD2 highlight substantial conformational flexibility of these SDR enzymes, which plays an important role in the establishment of catalytically productive complexes of the enzymes with their NADPH and phenolic substrates.
- Plant Biology Division, Samuel Roberts Noble Foundation, Ardmore, Oklahoma 73401.
Organizational Affiliation: