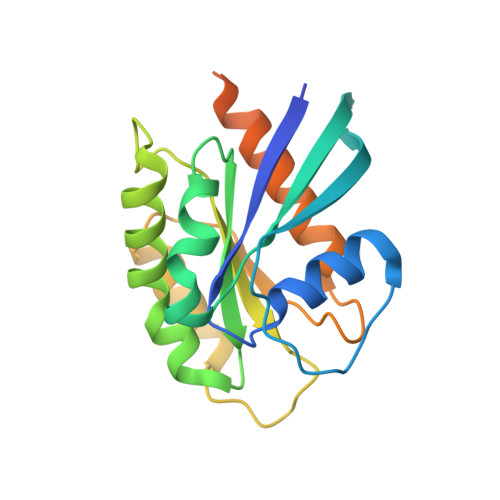
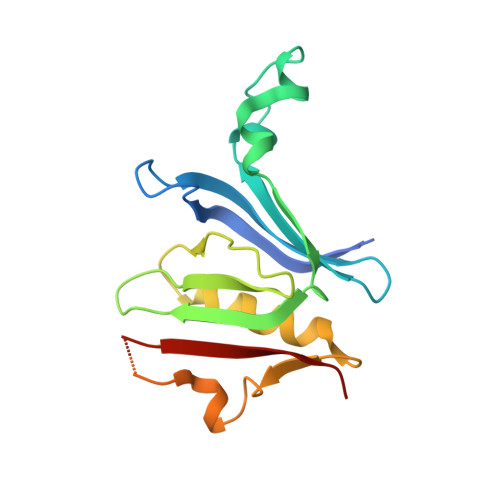
Crystal structure of the Rab9A-RUTBC2 RBD complex reveals the molecular basis for the binding specificity of Rab9A with RUTBC2.
Zhang, Z., Wang, S., Shen, T., Chen, J., Ding, J.(2014) Structure 22: 1408-1420
- PubMed: 25220469
- DOI: https://doi.org/10.1016/j.str.2014.08.005
- Primary Citation of Related Structures:
4QXA - PubMed Abstract:
Rab9 plays a vital role in regulating the transport of mannose 6-phosphate receptors from late endosomes to the trans-Golgi network through interactions with various effectors. Here, we report the crystal structure of GTP-bound Rab9A in complex with the Rab-binding domain (RBD) of the effector RUTBC2. RUTBC2 RBD assumes a pleckstrin homology domain fold that uses a binding site consisting of mainly β1 and the η1 insertion to interact with the switch and interswitch regions of Rab9A. The C-terminal hypervariable region of Rab9A is disordered and thus not required for RUTBC2 binding. The conformational plasticity of the switch and interswitch regions of Rab9A primarily determines the specificity for RUTBC2. Our biochemical and biological data confirm these findings and further show that Rab9B can bind to RUTBC2 probably in a similar manner as Rab9A. These results together reveal the molecular basis for the binding specificity of Rab9A with RUTBC2.
- State Key Laboratory of Molecular Biology, Institute of Biochemistry and Cell Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, 320 Yue-Yang Road, Shanghai 200031, China.
Organizational Affiliation: