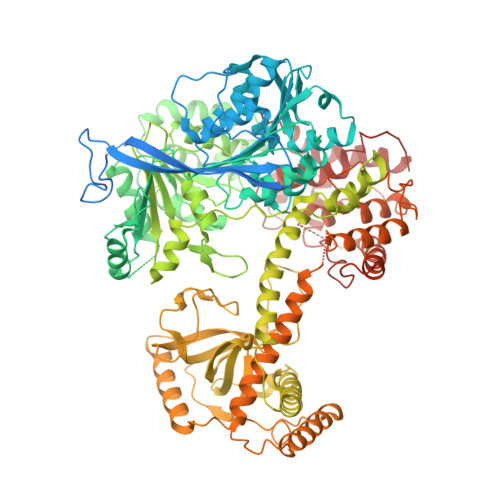
The Mtr4 ratchet helix and arch domain both function to promote RNA unwinding.
Taylor, L.L., Jackson, R.N., Rexhepaj, M., King, A.K., Lott, L.K., van Hoof, A., Johnson, S.J.(2014) Nucleic Acids Res 42: 13861-13872
- PubMed: 25414331
- DOI: https://doi.org/10.1093/nar/gku1208
- Primary Citation of Related Structures:
4QU4 - PubMed Abstract:
Mtr4 is a conserved Ski2-like RNA helicase and a subunit of the TRAMP complex that activates exosome-mediated 3'-5' turnover in nuclear RNA surveillance and processing pathways. Prominent features of the Mtr4 structure include a four-domain ring-like helicase core and a large arch domain that spans the core. The 'ratchet helix' is positioned to interact with RNA substrates as they move through the helicase. However, the contribution of the ratchet helix in Mtr4 activity is poorly understood. Here we show that strict conservation along the ratchet helix is particularly extensive for Ski2-like RNA helicases compared to related helicases. Mutation of residues along the ratchet helix alters in vitro activity in Mtr4 and TRAMP and causes slow growth phenotypes in vivo. We also identify a residue on the ratchet helix that influences Mtr4 affinity for polyadenylated substrates. Previous work indicated that deletion of the arch domain has minimal effect on Mtr4 unwinding activity. We now show that combining the arch deletion with ratchet helix mutations abolishes helicase activity and produces a lethal in vivo phenotype. These studies demonstrate that the ratchet helix modulates helicase activity and suggest that the arch domain plays a previously unrecognized role in unwinding substrates.
- Department of Chemistry and Biochemistry, Utah State University, Logan, UT 84322-0300, USA.
Organizational Affiliation: