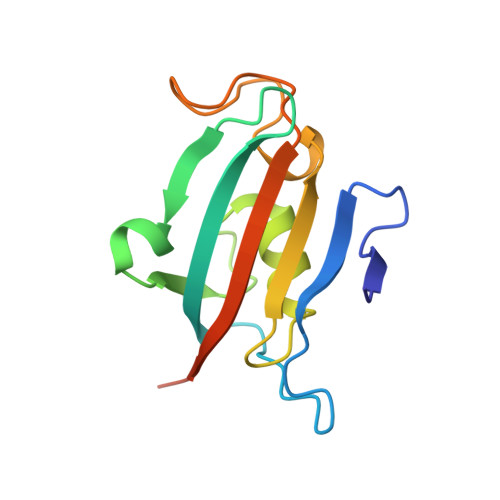
Two crystal structures of the FK506-binding domain of Plasmodium falciparum FKBP35 in complex with rapamycin at high resolution
Bianchin, A., Allemand, F., Bell, A., Chubb, A.J., Guichou, J.-F.(2015) Acta Crystallogr D Biol Crystallogr 71: 1319-1327
- PubMed: 26057671 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004715006239
- Primary Citation Related Structures:
4QT2, 4QT3 - PubMed Abstract:
Antimalarial chemotherapy continues to be challenging in view of the emergence of drug resistance, especially artemisinin resistance in Southeast Asia. It is critical that novel antimalarial drugs are identified that inhibit new targets with unexplored mechanisms of action. It has been demonstrated that the immunosuppressive drug rapamycin, which is currently in clinical use to prevent organ-transplant rejection, has antimalarial effects. The Plasmodium falciparum target protein is PfFKBP35, a unique immunophilin FK506-binding protein (FKBP). This protein family binds rapamycin, FK506 and other immunosuppressive and non-immunosuppressive macrolactones. Here, two crystallographic structures of rapamycin in complex with the FK506-binding domain of PfFKBP35 at high resolution, in both its oxidized and reduced forms, are reported. In comparison with the human FKBP12-rapamycin complex reported previously, the structures reveal differences in the β4-β6 segment that lines the rapamycin binding site. Structural differences between the Plasmodium protein and human hFKBP12 include the replacement of Cys106 and Ser109 by His87 and Ile90, respectively. The proximity of Cys106 to the bound rapamycin molecule (4-5 Å) suggests possible routes for the rational design of analogues of rapamycin with specific antiparasitic activity. Comparison of the structures with the PfFKBD-FK506 complex shows that both drugs interact with the same binding-site residues. These two new structures highlight the structural differences and the specific interactions that must be kept in consideration for the rational design of rapamycin analogues with antimalarial activity that specifically bind to PfFKBP35 without immunosuppressive effects.
- Conway Institute of Biomolecular and Biomedical Science, University College Dublin, Dublin, Ireland.
Organizational Affiliation: