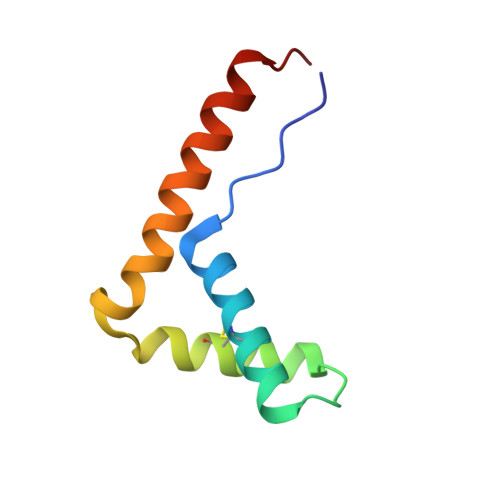
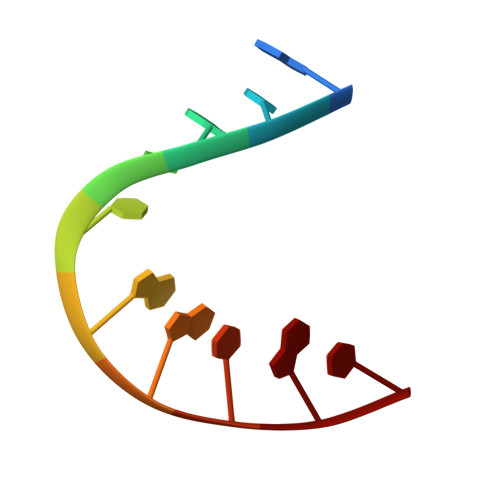
Two high-mobility group box domains act together to underwind and kink DNA.
Sanchez-Giraldo, R., Acosta-Reyes, F.J., Malarkey, C.S., Saperas, N., Churchill, M.E., Campos, J.L.(2015) Acta Crystallogr D Biol Crystallogr 71: 1423-1432
- PubMed: 26143914 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004715007452
- Primary Citation Related Structures:
4QR9 - PubMed Abstract:
High-mobility group protein 1 (HMGB1) is an essential and ubiquitous DNA architectural factor that influences a myriad of cellular processes. HMGB1 contains two DNA-binding domains, box A and box B, which have little sequence specificity but have remarkable abilities to underwind and bend DNA. Although HMGB1 box A is thought to be responsible for the majority of HMGB1-DNA interactions with pre-bent or kinked DNA, little is known about how it recognizes unmodified DNA. Here, the crystal structure of HMGB1 box A bound to an AT-rich DNA fragment is reported at a resolution of 2 Å. Two box A domains of HMGB1 collaborate in an unusual configuration in which the Phe37 residues of both domains stack together and intercalate the same CG base pair, generating highly kinked DNA. This represents a novel mode of DNA recognition for HMGB proteins and reveals a mechanism by which structure-specific HMG boxes kink linear DNA.
- Departament d'Enginyeria Quimica, Universitat Politecnica de Catalunya, 08028 Barcelona, Spain.
Organizational Affiliation: