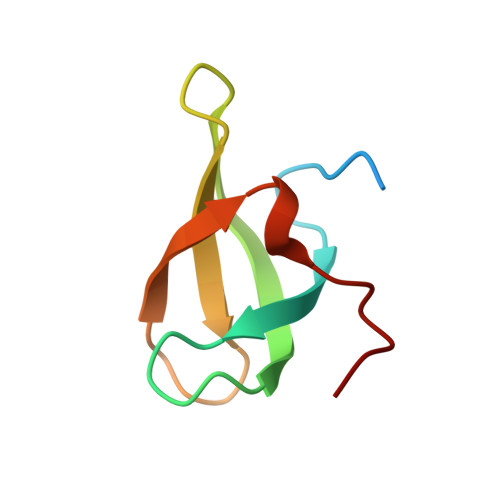
A small molecule antagonist of SMN disrupts the interaction between SMN and RNAP II.
Liu, Y., Iqbal, A., Li, W., Ni, Z., Wang, Y., Ramprasad, J., Abraham, K.J., Zhang, M., Zhao, D.Y., Qin, S., Loppnau, P., Jiang, H., Guo, X., Brown, P.J., Zhen, X., Xu, G., Mekhail, K., Ji, X., Bedford, M.T., Greenblatt, J.F., Min, J.(2022) Nat Commun 13: 5453-5453
- PubMed: 36114190 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-33229-5
- Primary Citation Related Structures:
4QQ6, 4QQD, 6V9T, 7W2P, 7W30 - PubMed Abstract:
Survival of motor neuron (SMN) functions in diverse biological pathways via recognition of symmetric dimethylarginine (Rme2s) on proteins by its Tudor domain, and deficiency of SMN leads to spinal muscular atrophy. Here we report a potent and selective antagonist with a 4-iminopyridine scaffold targeting the Tudor domain of SMN. Our structural and mutagenesis studies indicate that both the aromatic ring and imino groups of compound 1 contribute to its selective binding to SMN. Various on-target engagement assays support that compound 1 specifically recognizes SMN in a cellular context and prevents the interaction of SMN with the R1810me2s of RNA polymerase II subunit POLR2A, resulting in transcription termination and R-loop accumulation mimicking SMN depletion. Thus, in addition to the antisense, RNAi and CRISPR/Cas9 techniques, potent SMN antagonists could be used as an efficient tool to understand the biological functions of SMN.
- Jiangsu Key Laboratory of Neuropsychiatric Diseases and College of Pharmaceutical Sciences, Soochow University, Suzhou, Jiangsu, China. ylliu18@suda.edu.cn.
Organizational Affiliation: