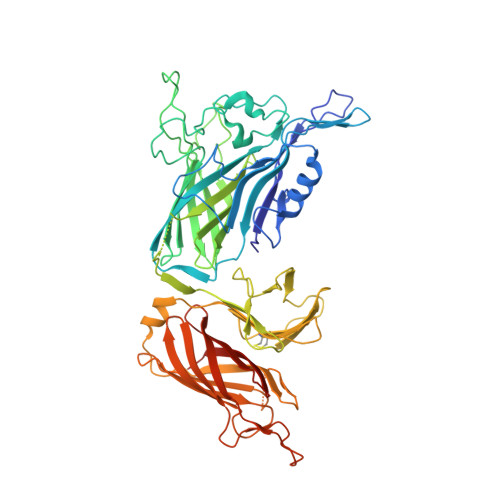
Patterns of structural and sequence variation within isotype lineages of the Neisseria meningitidis transferrin receptor system.
Adamiak, P., Calmettes, C., Moraes, T.F., Schryvers, A.B.(2015) Microbiologyopen 4: 491-504
- PubMed: 25800619
- DOI: https://doi.org/10.1002/mbo3.254
- Primary Citation of Related Structures:
4QQ1 - PubMed Abstract:
Neisseria meningitidis inhabits the human upper respiratory tract and is an important cause of sepsis and meningitis. A surface receptor comprised of transferrin-binding proteins A and B (TbpA and TbpB), is responsible for acquiring iron from host transferrin. Sequence and immunological diversity divides TbpBs into two distinct lineages; isotype I and isotype II. Two representative isotype I and II strains, B16B6 and M982, differ in their dependence on TbpB for in vitro growth on exogenous transferrin. The crystal structure of TbpB and a structural model for TbpA from the representative isotype I N. meningitidis strain B16B6 were obtained. The structures were integrated with a comprehensive analysis of the sequence diversity of these proteins to probe for potential functional differences. A distinct isotype I TbpA was identified that co-varied with TbpB and lacked sequence in the region for the loop 3 α-helix that is proposed to be involved in iron removal from transferrin. The tightly associated isotype I TbpBs had a distinct anchor peptide region, a distinct, smaller linker region between the lobes and lacked the large loops in the isotype II C-lobe. Sequences of the intact TbpB, the TbpB N-lobe, the TbpB C-lobe, and TbpA were subjected to phylogenetic analyses. The phylogenetic clustering of TbpA and the TbpB C-lobe were similar with two main branches comprising the isotype 1 and isotype 2 TbpBs, possibly suggesting an association between TbpA and the TbpB C-lobe. The intact TbpB and TbpB N-lobe had 4 main branches, one consisting of the isotype 1 TbpBs. One isotype 2 TbpB cluster appeared to consist of isotype 1 N-lobe sequences and isotype 2 C-lobe sequences, indicating the swapping of N-lobes and C-lobes. Our findings should inform future studies on the interaction between TbpB and TbpA and the process of iron acquisition.
- Department of Microbiology, Immunology and Infectious Diseases, University of Calgary, Calgary, Alberta, T2N 4N1, Canada.
Organizational Affiliation: