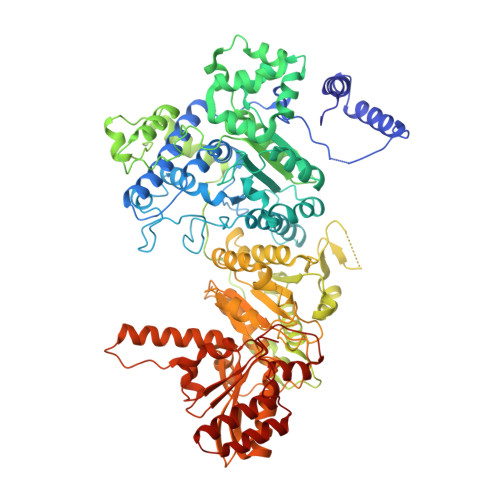
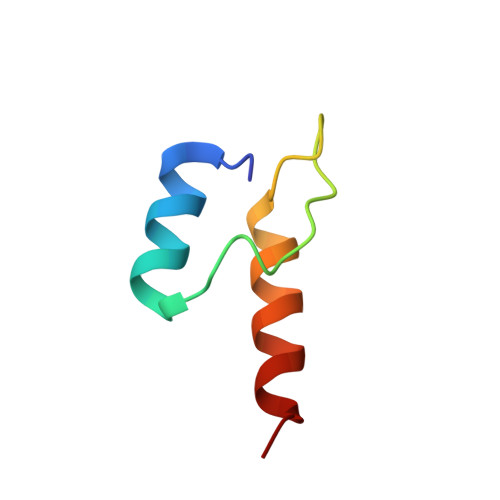
Novel Binding Motif and New Flexibility Revealed by Structural Analyses of a Pyruvate Dehydrogenase-Dihydrolipoyl Acetyltransferase Subcomplex from the Escherichia coli Pyruvate Dehydrogenase Multienzyme Complex.
Arjunan, P., Wang, J., Nemeria, N.S., Reynolds, S., Brown, I., Chandrasekhar, K., Calero, G., Jordan, F., Furey, W.(2014) J Biological Chem 289: 30161-30176
- PubMed: 25210042 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.592915
- Primary Citation Related Structures:
4QOY - PubMed Abstract:
The Escherichia coli pyruvate dehydrogenase multienzyme complex contains multiple copies of three enzymatic components, E1p, E2p, and E3, that sequentially carry out distinct steps in the overall reaction converting pyruvate to acetyl-CoA. Efficient functioning requires the enzymatic components to assemble into a large complex, the integrity of which is maintained by tethering of the displaced, peripheral E1p and E3 components to the E2p core through non-covalent binding. We here report the crystal structure of a subcomplex between E1p and an E2p didomain containing a hybrid lipoyl domain along with the peripheral subunit-binding domain responsible for tethering to the core. In the structure, a region at the N terminus of each subunit in the E1p homodimer previously unseen due to crystallographic disorder was observed, revealing a new folding motif involved in E1p-E2p didomain interactions, and an additional, unexpected, flexibility was discovered in the E1p-E2p didomain subcomplex, both of which probably have consequences in the overall multienzyme complex assembly. This represents the first structure of an E1p-E2p didomain subcomplex involving a homodimeric E1p, and the results may be applicable to a large range of complexes with homodimeric E1 components. Results of HD exchange mass spectrometric experiments using the intact, wild type 3-lipoyl E2p and E1p are consistent with the crystallographic data obtained from the E1p-E2p didomain subcomplex as well as with other biochemical and NMR data reported from our groups, confirming that our findings are applicable to the entire E1p-E2p assembly.
- From the Departments of Pharmacology and Chemical Biology and.
Organizational Affiliation: