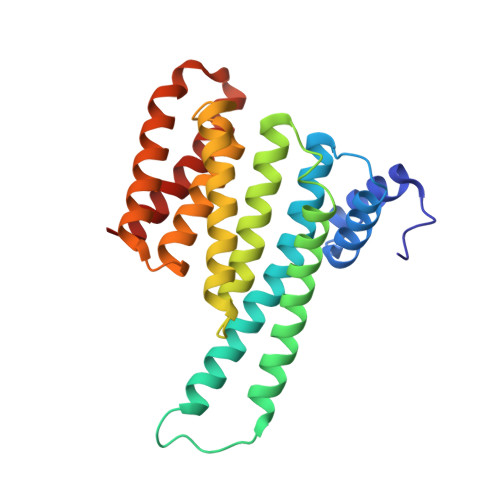
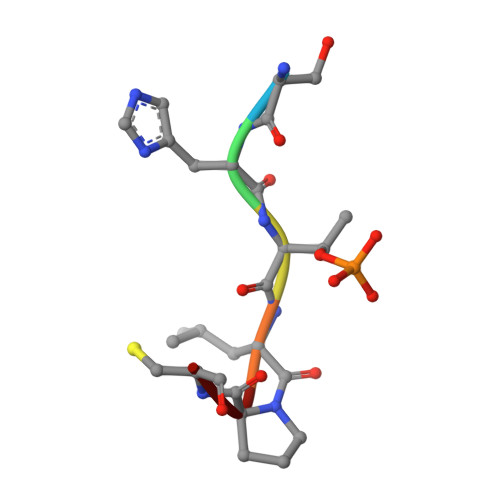
A novel phospho-switch in the linker region of the snail zinc finger protein which regulates 14-3-3 association, DNA binding and epithelial-mesenchymal differentiation
Liu, Y., Prokop, J.W., Peng, H., Karar, J., White, D., Hou, Z., Qin, Y.S., Bier, D., Ottmann, C., Waxman, S., Halazonetis, T.D., Rauscher III, F.J.To be published.