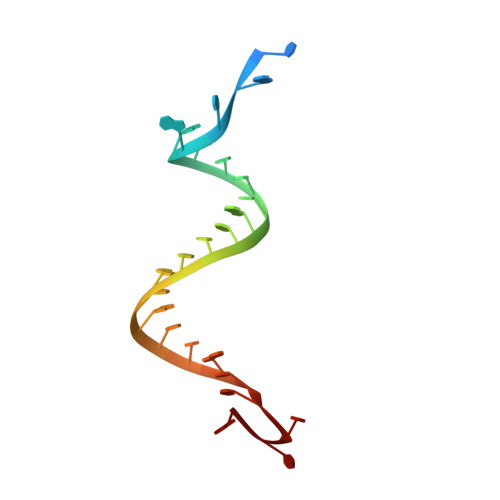
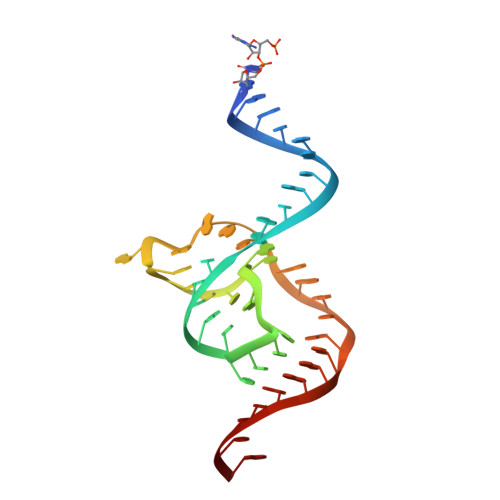
Structural basis for the fast self-cleavage reaction catalyzed by the twister ribozyme.
Eiler, D., Wang, J., Steitz, T.A.(2014) Proc Natl Acad Sci U S A 111: 13028-13033
- PubMed: 25157168 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1414571111
- Primary Citation Related Structures:
4QJD, 4QJH - PubMed Abstract:
Twister is a recently discovered RNA motif that is estimated to have one of the fastest known catalytic rates of any naturally occurring small self-cleaving ribozyme. We determined the 4.1-Å resolution crystal structure of a twister sequence from an organism that has not been cultured in isolation, and it shows an ordered scissile phosphate and nucleotide 5' to the cleavage site. A second crystal structure of twister from Orzyza sativa determined at 3.1-Å resolution exhibits a disordered scissile phosphate and nucleotide 5' to the cleavage site. The core of twister is stabilized by base pairing, a large network of stacking interactions, and two pseudoknots. We observe three nucleotides that appear to mediate catalysis: a guanosine that we propose deprotonates the 2'-hydroxyl of the nucleotide 5' to the cleavage site and a conserved adenosine. We suggest the adenosine neutralizes the negative charge on a nonbridging phosphate oxygen atom at the cleavage site. The active site also positions the labile linkage for in-line nucleophilic attack, and thus twister appears to simultaneously use three strategies proposed for small self-cleaving ribozymes. The twister crystal structures (i) show its global structure, (ii) demonstrate the significance of the double pseudoknot fold, (iii) provide a possible hypothesis for enhanced catalysis, and (iv) illuminate the roles of all 10 highly conserved nucleotides of twister that participate in the formation of its small and stable catalytic pocket.
- Department of Molecular Biochemistry and Biophysics and.
Organizational Affiliation: