Structural transition of a protein nanocage as a function of pH and salt concentration.
Lai, Y.-T., Yeates, T.O.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Non-haem bromoperoxidase BPO-A2, Matrix protein 1 chimera | 456 | Kitasatospora aureofaciens, Influenza A virus This entity is chimeric | Mutation(s): 3 Gene Names: bpoA2, M EC: 1.11.1 | 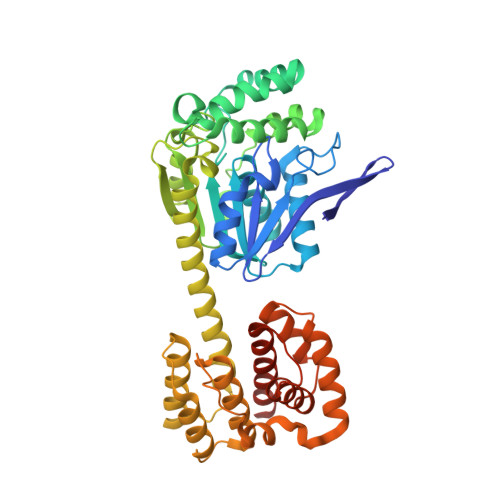 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Groups | P29715P03485 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 123.51 | α = 90 |
| b = 165.52 | β = 90 |
| c = 167.67 | γ = 90 |
| Software Name | Purpose |
|---|---|
| XSCALE | data scaling |
| PHASER | phasing |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |